ImJoy Team
84 posts

ImJoy Team
@ImJoyTeam
ImJoy(https://t.co/PpDSqaHaEy): a platform for deploying deep learning tools, built with PWA, WebAssembly, Python, Jupyter and more.






Do you want to learn how to use applications such as #ZeroCostDL4Mic, @ilastik_team, @ImJoyTeam, the @BioimageA, and @CellProfiler? 💻 Microscopy data analysis: machine learning and the #BioImageArchive 🔗 Apply now: ebi.ac.uk/training/event…



Seems this week is full of celebrations 🥳 our contribution to make DL more accessible has been published in @naturemethods and is already available 🔥🔥🔥 "DeepImageJ: A user-friendly environment to run deep learning models in ImageJ" nature.com/articles/s4159…



Fully reproducible biological experiments? Here we go: A "SMART" workflow using sharable @ProjectJupyter Notebooks including @opentrons robots, @ImJoyTeam image processing and an @OpenFlexure-controlled @OpenUc2 fluorescence high-throughput microscope. bit.ly/3j9pOtF 1/n






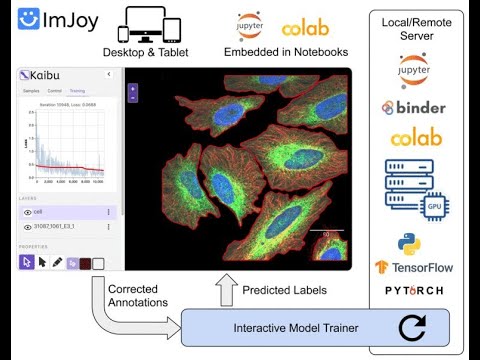
New #ZeroCostDL4Mic release: Kaibu/ImJoy's Interactive Segmentation! Annotate and Train simultaneously #HumanInTheLoop #DeepLearning colab.research.google.com/github/Henriqu… Many thanks to @weioyang and @ImJoyTeam for the support and this: #ref-15" target="_blank" rel="nofollow noopener">f1000research.com/articles/10-14…
Data courtesy of @Roubinet31 !
I'm so excited by the possibilities of connecting up open hardware and software. @openflexure + @ImJoyTeam, brought together by @OpenUc2, mean we can build ImageJ into our microscope interface! (gitlab.com/openflexure/op…)

Hi all! I'm glad to announce the new release of #deepImageJ. Together with @carlosg91018370, @gomez_mariscal and @DanielSage, we've got an evolved #deepImageJ with exciting new features 🤓 -#BioImageModelZoo -#PyTorch -#3D, #classification, #detection -#gpu -#java proccessings


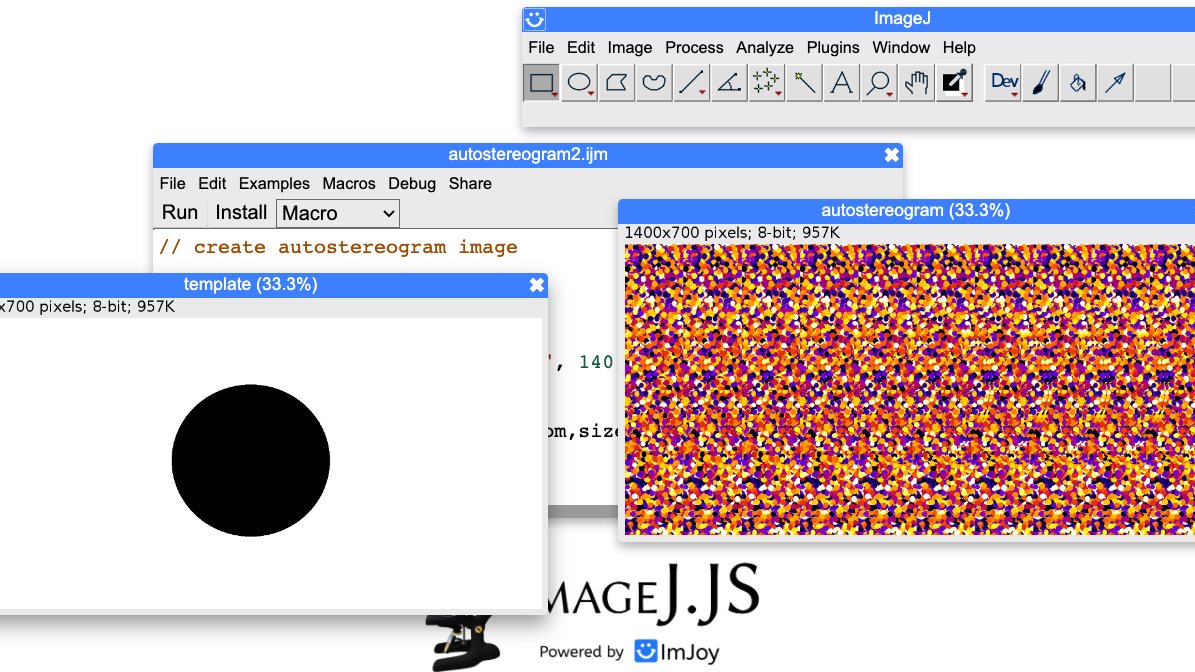
Instead of using a closed source online autostereogram maker, I decided to create one from scratch in ImageJ. github.com/mutterer/weird…

