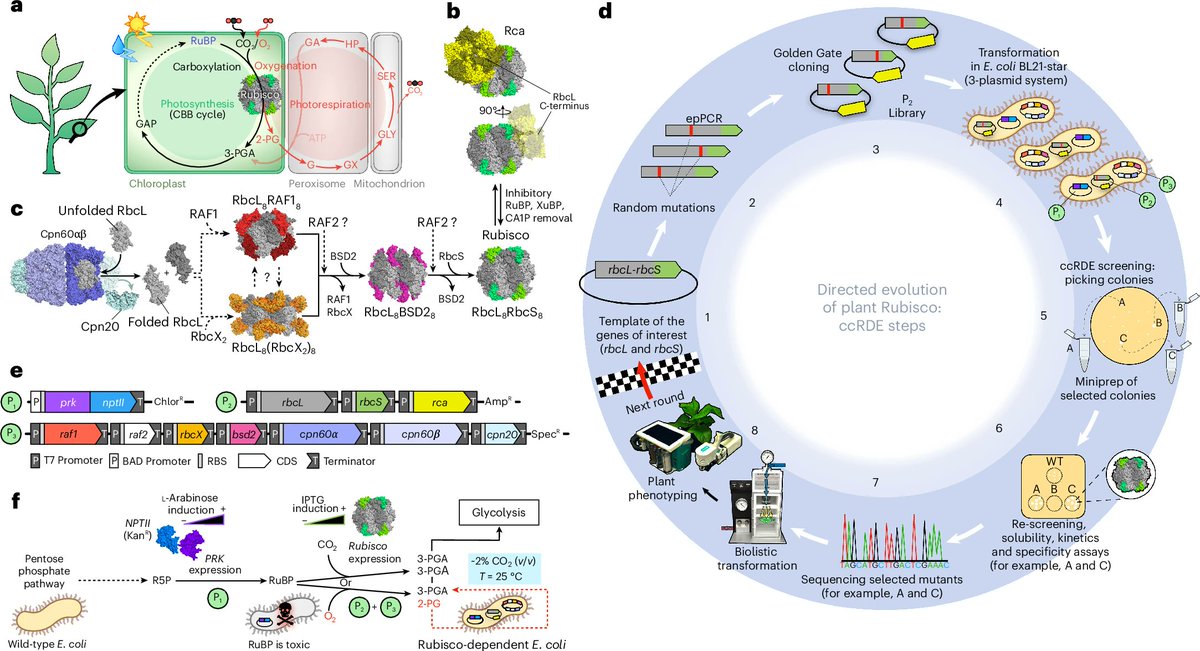
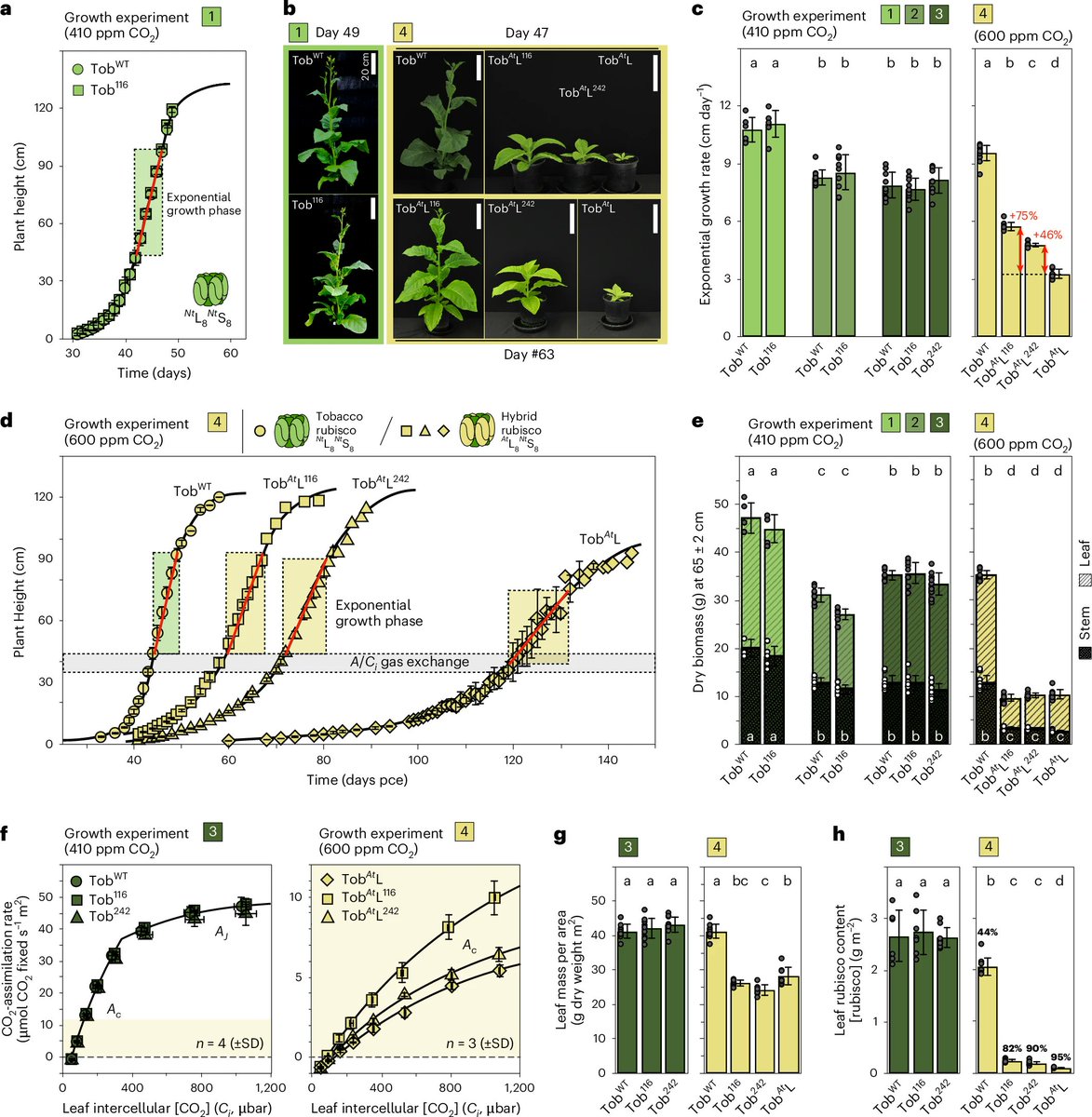
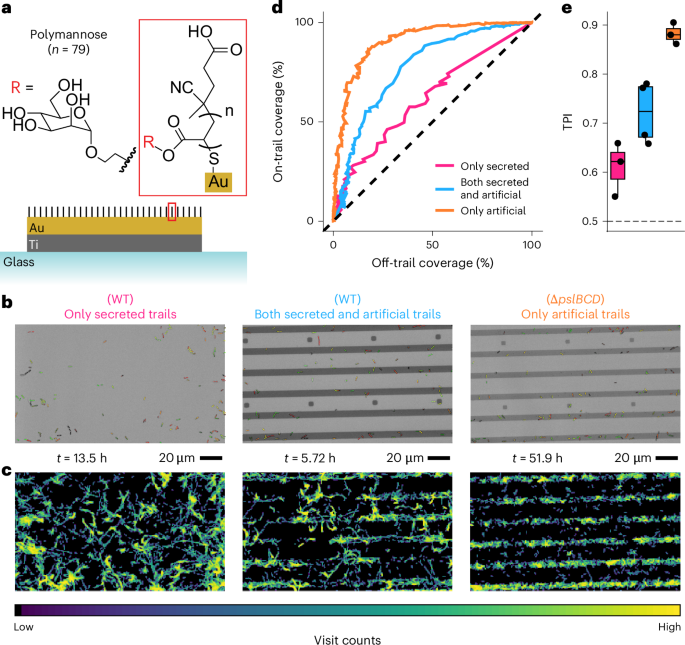
The complex developmental mechanisms of nucleus-forming jumbo phages
Mozumdar D, Agard DA, Bondy-Denomy J
Curr Opin Microbiol. 88:102676, doi: 10.1016/j.mib.2025.102676
pubmed.ncbi.nlm.nih.gov/41016252/
sciencedirect.com/science/articl…
English
JBacteriology Eds
565 posts

@JBacteriology
Tweets from the Editors of the Journal of Bacteriology. Revived on August 4, 2022