Justus Großmann retweetledi

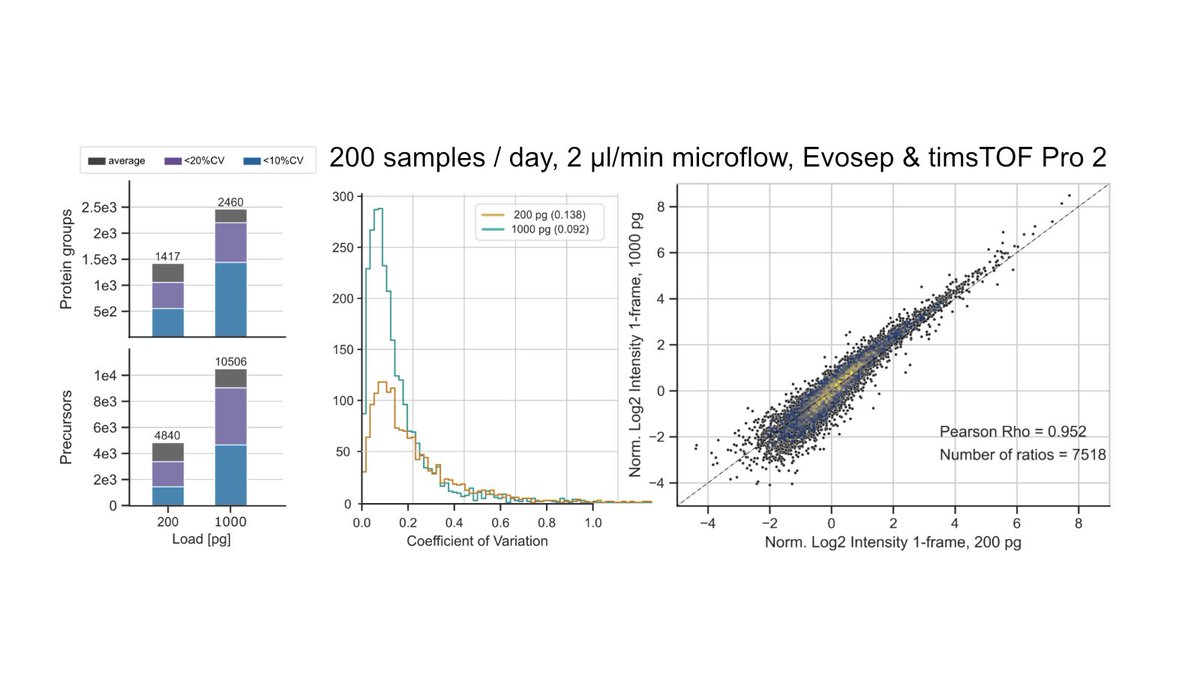
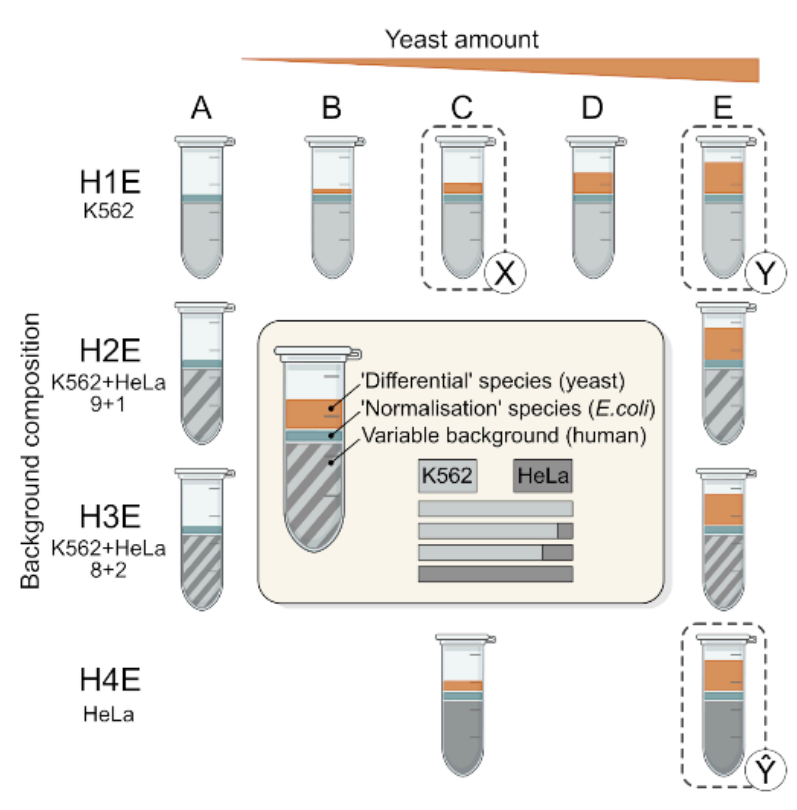
We acquired a large-scale mixed-species benchmark, with variable background, to comprehensively assess quantitative accuracy of proteomics.
Key features:
- 192 runs, 0.75ng - 15ng of variable human cell line backgroud.
- Can dissect the impact of both random and systematic errors.
- Can test how quantitation algorithms scale with experiment size and sample heterogeneity.
Our insights based on the data: biorxiv.org/content/10.648….
PRIDE repo will be made public in the next days.
English