Kevin Bijan Givechian retweetledi

Yale medicine feature on our Immunostruct model medicine.yale.edu/news-article/u… !
English
Kevin Bijan Givechian
417 posts

@KevinGivechian
MD-PhD student @Yale | Co-founder @AscentBio | Exploring cancer bio, ML, protein engineering, & immunotherapy with @KrishnaswamyLab and @VirusesImmunity






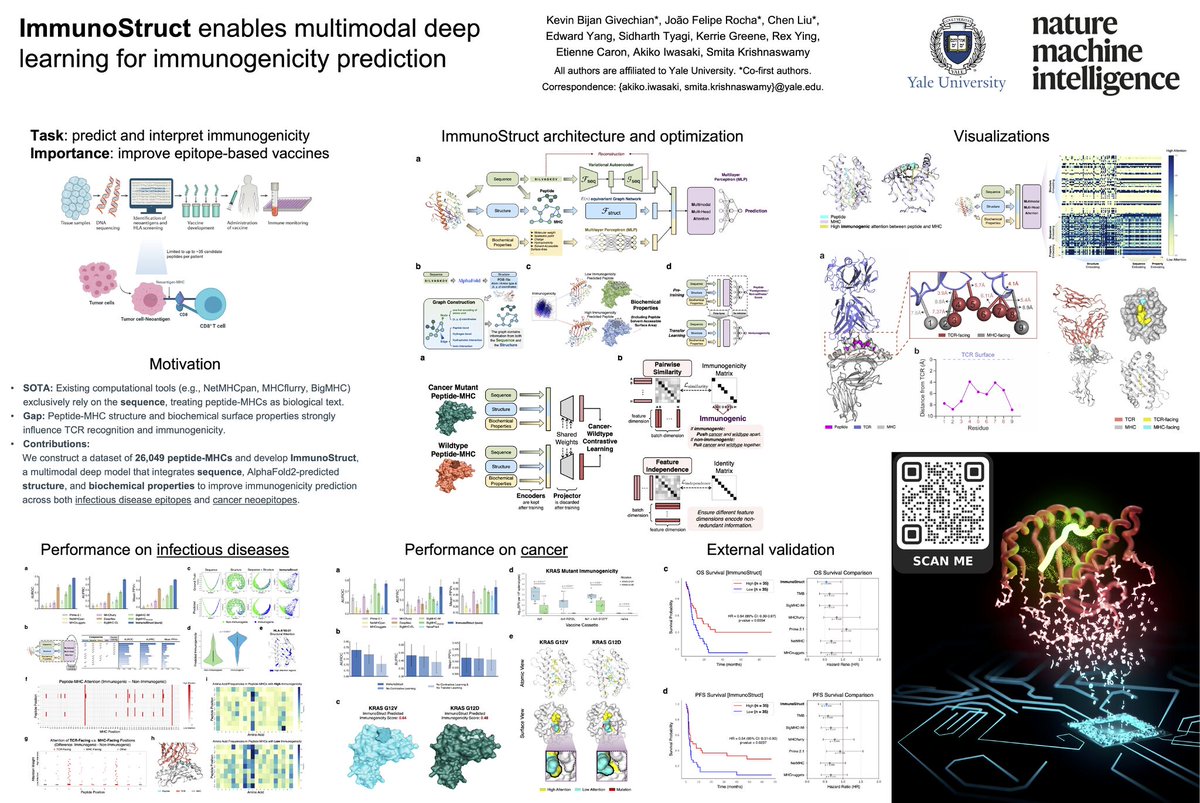





(1/n) Just in time for New Years! #ImmunoStruct, our multimodal model that predicts class I peptide-MHC immunogenicity is out at @NatMachIntell ! nature.com/articles/s4225…













