Sabitlenmiş Tweet
Kuzuyama_Lab
203 posts


Kuzuyama_Lab
@Kuzuyama_Lab
Synthetic Biology Lab. at The University of Tokyo. We aim to create new useful compounds and understand complex interactions between microorganisms in nature.
Tokyo, Japan Katılım Şubat 2021
12 Takip Edilen306 Takipçiler
Kuzuyama_Lab retweetledi

The Wakimoto (A03) and Yoshimura (A01) groups show that bacterial membrane vesicles (MVs) induce silent metabolite production, leading to the discovery of four unique N-alkylpyridinium alkaloids, stigmatellamines. pubs.acs.org/doi/full/10.10…
English
Kuzuyama_Lab retweetledi

The Wakimoto group (A03), in collaboration with Prof. J. Piel (ETH), reveals the long-elusive mechanism of nitrile formation in calyculin A through comparative analysis of two biosynthetic gene clusters from distinct producers. doi.org/10.1002/anie.3…
English
Kuzuyama_Lab retweetledi

The Kudo (A03) and Awakawa (A01) groups, in collaboration with Dr. Shin-ya, noted five-step editing of a type I polyketide synthase with minimal loss of production titre. The post-AT cut site would be the first choice for PKS designers. #PKS @NatureComms doi.org/10.1038/s41467…
English
Kuzuyama_Lab retweetledi

Hijacking tRNA chemistry to redesign antibiotics. The Maruyama group (A1), with the Ogasawara (A3) and Goto (A3) groups, uncovered a versatile enzyme generating new streptothricin derivatives and expanding antibiotic chemical diversity. @J_A_C_S doi.org/10.1021/jacs.6…
English
Kuzuyama_Lab retweetledi

The Umemura group (A02), in collaboration with the Kuzuyama group (A01), reports a transformer-based platform that predicts and generates functional domains within biosynthetic gene clusters via unsupervised learning of domain organization in genomes. journals.plos.org/ploscompbiol/a…
English

Lab article: Our work on the biosynthesis of the neuroprotective natural product kaitocephalin is now published in Angewandte Chemie. Genomic, enzymatic, and isotope-labeling analyses clarify key steps of its biosynthesis. onlinelibrary.wiley.com/doi/10.1002/an…
English
Kuzuyama_Lab retweetledi

The Nakano group (A02), in collaboration with the Chisuga group (A02), designed a high-fidelity sortase E with an activity trade-off. This enzyme can be applied to the synthesis of antibody conjugates. pubs.acs.org/doi/10.1021/ac…
English
Kuzuyama_Lab retweetledi

The Kato group (A03) reports a PCA-based database mining strategy to discover novel bacterial carbene transferases capable of stereodivergent cyclopropanation, highlighting a data-driven route to biocatalysts for abiotic transformations. doi.org/10.1002/anie.2…
English
Kuzuyama_Lab retweetledi

The Katsuyama Group (A03) revelaed that ATP-dependent diazotase is involved in the biosynthesis of nitrous acid-dependent aromatic amination to produce 2,4-diamino-3-hydroxybenzoic acid. …mistry-europe.onlinelibrary.wiley.com/doi/10.1002/cb…
English
Kuzuyama_Lab retweetledi

The Shoji group (A03), in collaboration with the Nakano group (A02), computationally redesigned the small heme protein HasA, achieving a +17.5 °C increase in thermostability while retaining metalloporphyrin binding, enabling robust HasA use. doi.org/10.1093/chemle…
English
Kuzuyama_Lab retweetledi

🎉 Our members presented at a symposium at #Pacifichem2025, sharing our research with the world 🌏

English
Kuzuyama_Lab retweetledi

第七回公開シンポジウムの参加登録を開始しました!bio-4cast.skr.jp/251208-2/
2026年2月13-14日に理化学研究所(和光市)鈴木梅太郎ホールで開催いたします。奮ってご参加ください!プログラムはこちら→bio-4cast.skr.jp/251119-2/
日本語
Kuzuyama_Lab retweetledi

Kuzuyama (A01), Wakimoto (A03), together with J. Piel (ETH) and H. Takeyama (Waseda Univ.), unveiled new ‘Entotheonella’ symbionts in marine sponges that possess diverse NRPS, PKS, RiPP, terpene synthase genes, highlighting remarkable enzymatic diversity. nature.com/articles/s4158…
English
Kuzuyama_Lab retweetledi

The Wakimoto group (A03) repurposed promiscuous head-to-tail NRP cyclases into head-to-side chain mode through substrate engineering—establishing a modular chemoenzymatic platform for lariat lipopeptides. nature.com/articles/s4155… @NatureChemistry
English
Kuzuyama_Lab retweetledi

The Ishitani (A02) and Terada (A02) groups developed an improved protein–ligand complex structure prediction based on Boltz-1. This software perfectly reproduces the chirality specified in the input chemical structure. Available at: github.com/cddlab/colabfo…
pubs.acs.org/doi/10.1021/ac…
English

Lab events: Mission accomplished! UTokyo gacha completed! Cheers to the lab crew for joining the fun!!
u-tokyo.ac.jp/focus/ja/artic…

English
Kuzuyama_Lab retweetledi

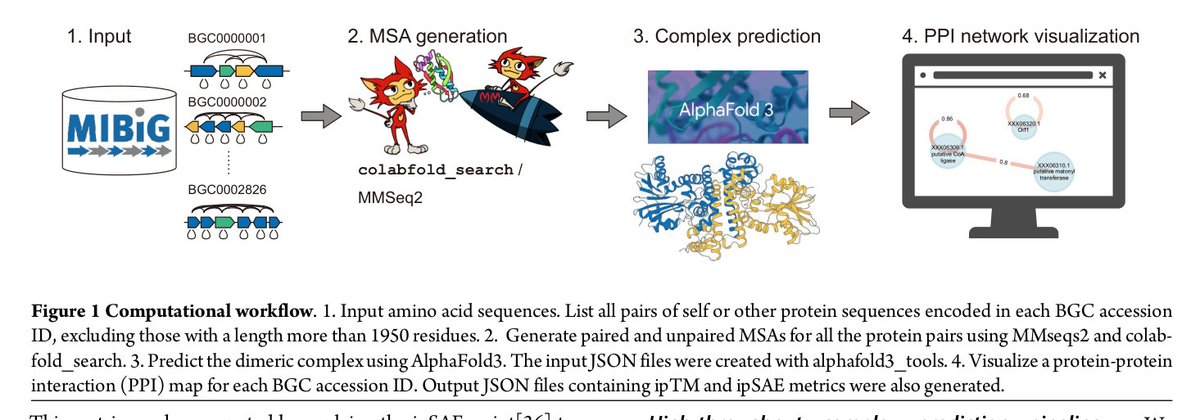
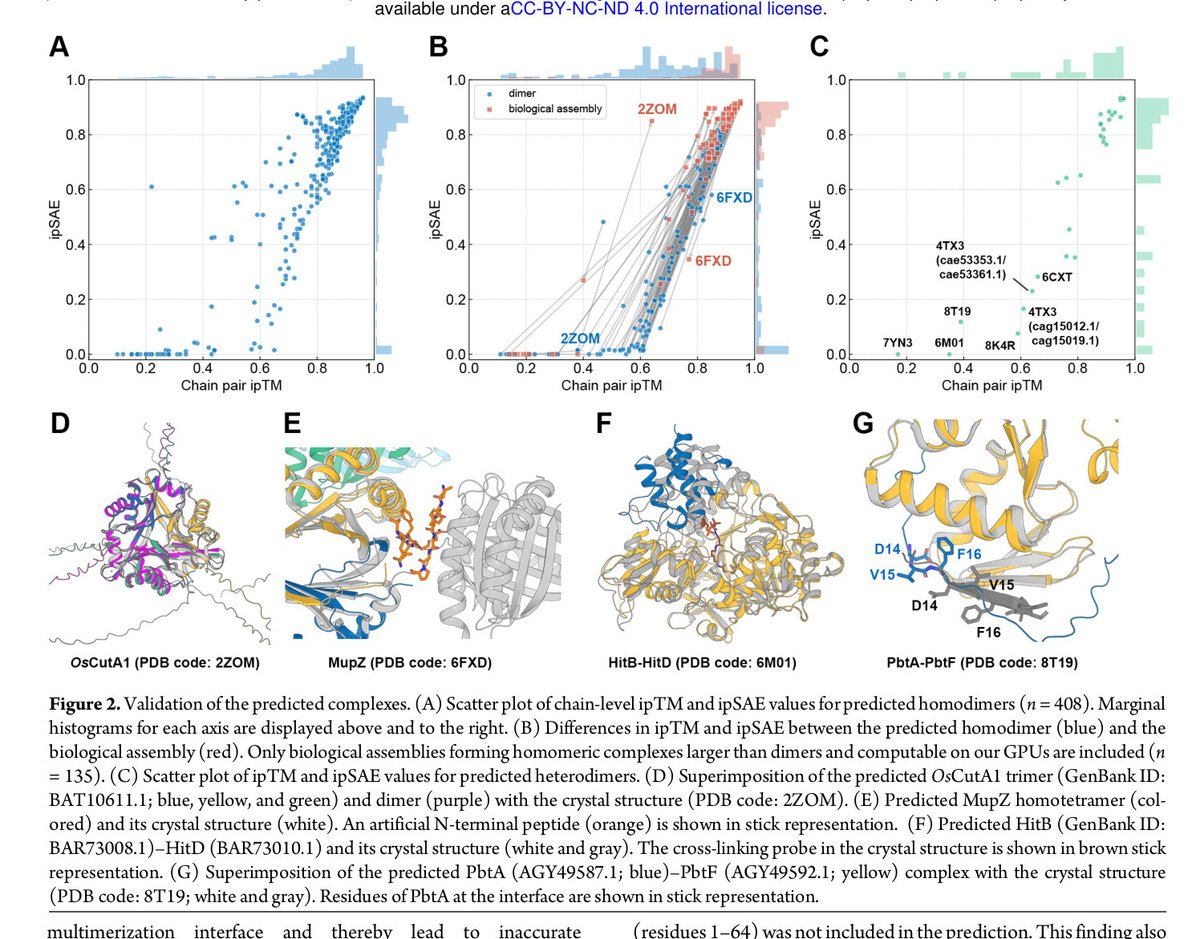
The Terada (A02) group, with Ishitani (A02), Wakimoto (A03), Minami (A03), Katsuyama (A03), and Kuzuyama (A01), predicted 487,828 complexes for known BGCs, identifying 15,438 heteromeric interactions.
biorxiv.org/content/10.110…
#Biosynthesis #ProteinComplexes #Breakthrough


English
Kuzuyama_Lab retweetledi

The Kuzuyama group unveiled key steps in the biosynthesis of kaitocephalin, a neuroprotective agent discovered 30 years ago here. Identifying kpb BGC pried open the long-locked door toward full elucidation. doi.org/10.1101/2025.1…
#Breakthrough #NaturalProducts #DrugDiscovery
English

Kaitocephalin, a neuroprotective metabolite discovered here 30 years ago, remained a biosynthetic mystery. We unveiled its kpb BGC & pathway steps, prying open the door to full elucidation and applications.
🔗 doi.org/10.1101/2025.1…
#Breakthrough #NaturalProducts #DrugDiscovery
English
