Li-Hsin Chang
307 posts


Li-Hsin Chang
@Lihsin0507
Postdoc @jdavieslab @MRC_WIMM @UniofOxford - 3D genome organization, TADs, CTCF, genome editing & Nanopore sequencing
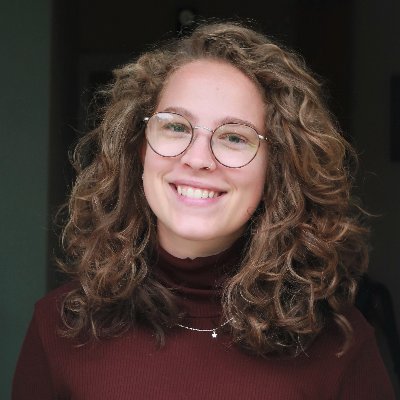






Congratulations to our students @grace_meaker and @IsaacSWalton, who both won awards at the Oxford MRC Doctoral Training Partnership Symposium (@Oxford_MRC_DTP) earlier today. Grace won the first prize in the student short talks, with her piece on 'The Novel Role of Runx Factors: Regulating Haematopoietic Stem Cell Expansion'. Isaac won the Judge’s Award in the 3-Minute Research Competition. Well done both!















Today, we - @stark_lab and @Eileen_Furlong lab (@EMBLheidelberg) - report the design of 40 synthetic enhancers for 5 tissue types in fruit fly embryos, in vivo. With only a few hundred enhancers per tissue for training, we adopt a stepping-stone transfer-learning approach: (5/N)