
Lucas Czech
95 posts

Lucas Czech
@LucCzech
Postdoc in bioinformatics 🌱 and computer scientist 🤖 working on interdisciplinary ways to save the planet 🌍














Another episode of "have you ever fought with software X?" Current tools for pool-seq in Evolve+Reseq data are too slow or have other problems (🧵) @LucCzech wrote from scratch an easy to use C++ tool for pool-seq + comp popgen stats w @Spence_Jeffrey_ arxiv.org/abs/2306.11622








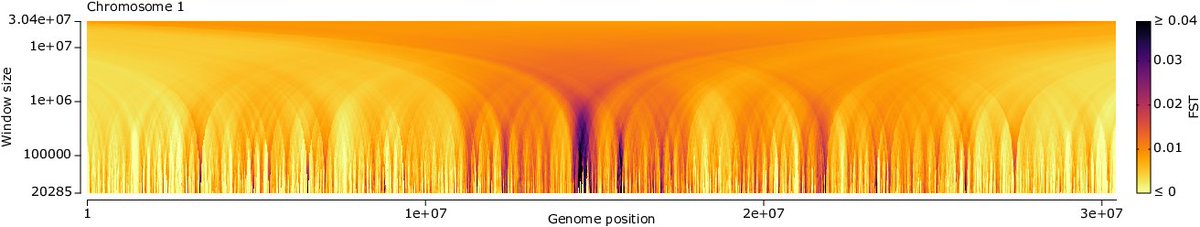

Another episode of "have you ever fought with software X?" Current tools for pool-seq in Evolve+Reseq data are too slow or have other problems (🧵) @LucCzech wrote from scratch an easy to use C++ tool for pool-seq + comp popgen stats w @Spence_Jeffrey_ arxiv.org/abs/2306.11622



The #MOILAB will be at #evolution2023 presenting in diverse topics: experimental evolution and popgen, ecology and local adaptation, to conservation biology, and biogeography! evolutionmeetings.org @CarnegiePlants @CarnegieEcolo @Stanford @HHMINEWS moilab.science/team






