Mathilde Hutin
222 posts


Mathilde Hutin retweetledi

Evolutionary and epidemiological insights from historical and modern genomes of Xanthomonas oryzae pv. oryzicola, the causal agent of bacterial leaf streak of rice | Molecular Plant-Microbe Interactions®
apsjournals.apsnet.org/doi/10.1094/MP…
English
Mathilde Hutin retweetledi

How to celebrate the 20th anniversary of the first #Xanthomonas genomes? Writing an essay could be the solution. peercommunityjournal.org/item/10_24072_…
We thank @AgenceRecherche and @COSTprogramme. @NicolasWGChen @Mathilde_Hutin @alperezqui @Perr_Po @RieuxAdrien @boris_szurek @PHIM_research

English
Mathilde Hutin retweetledi

#LeMomentPresse📻@RFI |👉#Madagascar: la bactériose vasculaire, maladie contagieuse sur le #riz, inquiète les chercheurs
🔴Les pertes de rendement sur les parcelles infectées peuvent aller jusqu’à 70%. Itw de @Mathilde_Hutin, chercheuse à l'#IRD qui étudie les souches contaminées
RFI@RFI
Madagascar: la bactériose vasculaire, maladie contagieuse sur le riz, inquiète les chercheurs rfi.my/AHGx.x
Français
Mathilde Hutin retweetledi

@ZoeDubrow @icpp2023 Owwww Zoe, you have done it 👏🏻💪! So happy to know that we’ll see each other in few days !
English

270km, 4 days, and lots of incredible views later, these two plant pathologists have made it to Lyon! Excited for @icpp2023!!


English
Mathilde Hutin retweetledi

[#phimmediation] 🗣️#DidierTharreau vous parle de #riz et vous explique pourquoi repenser la #riziculture est une nécessité 🌾🌾🌾@cirad #Okoo @ceciledjunga @MaxBirdOfficiel @PrFeuillage
france.tv/france-4/c-est…
Français
Mathilde Hutin retweetledi
Genetic Structure and TALome Analysis Highlight a High Level of Diversity in Burkinabe Xanthomonas Oryzae pv. oryzae Populations sco.lt/6E3U1o

English
Mathilde Hutin retweetledi

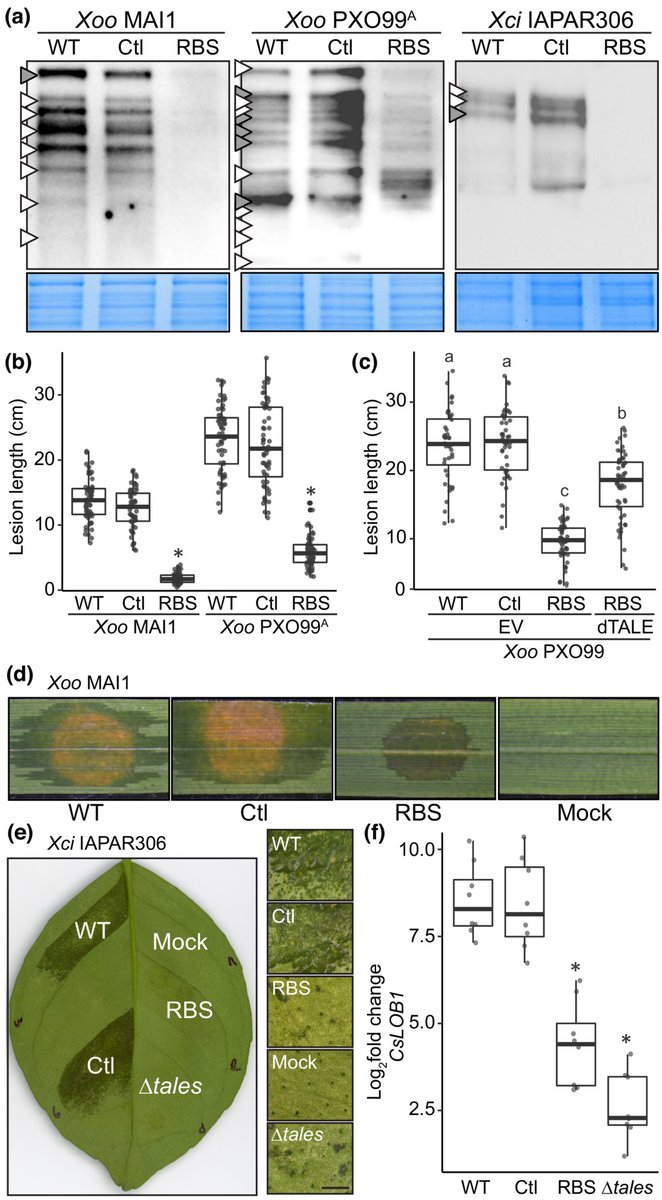
Multigene family silencing through CRISPR interference in Xanthomonas
Zárate-Chaves et al. @boris_szurek @koebnik_ralf @adrianabernalg
📖ow.ly/3QN250Nqtuy
English
Mathilde Hutin retweetledi

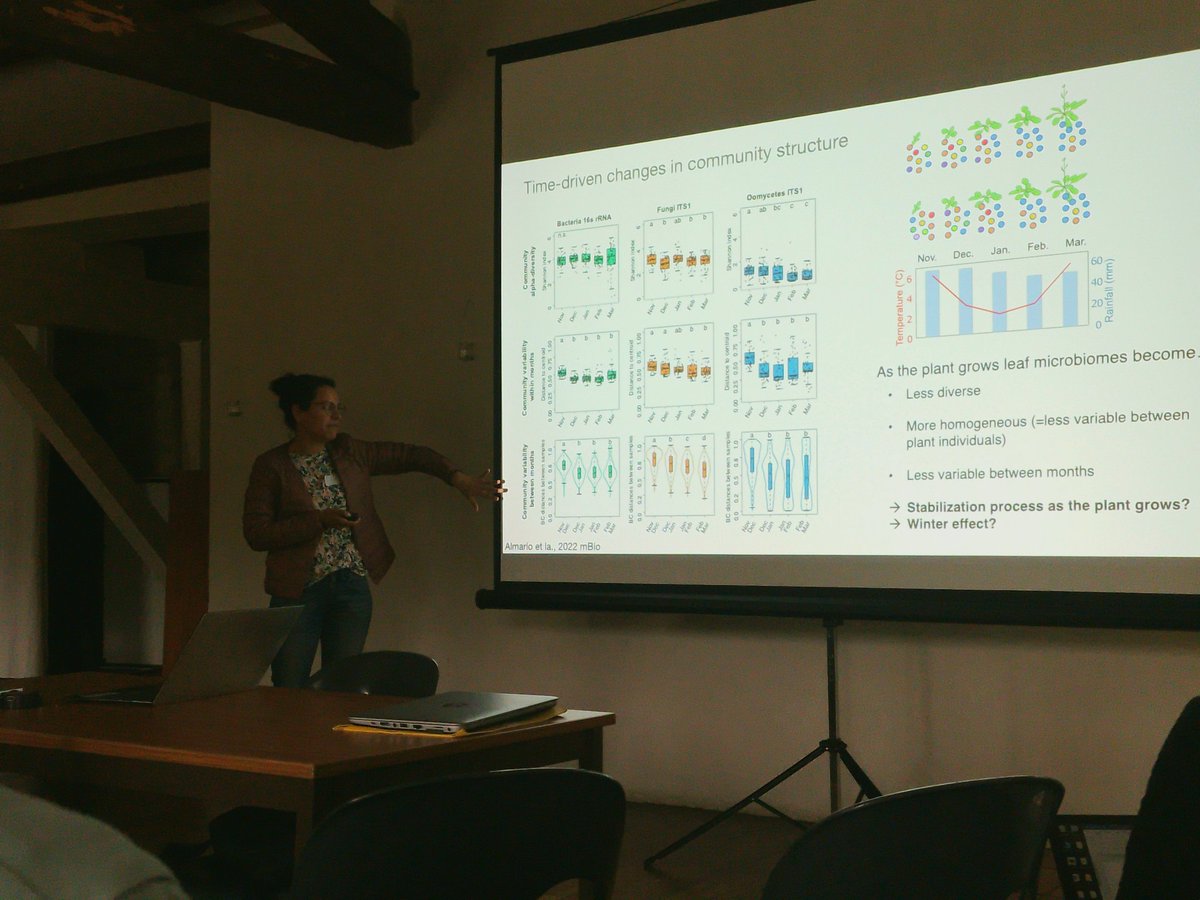
[#Meeting] Le colloque #MoDiP2022 "Molecular dialogue in plant biotic interactions" commence avec les présentations de @julianaalma, #SaraDjouadou, @LWaleryszak et #NatashaHortaAraújo pour la 1ère session, à l'auberge du Cèdre.



Français
Mathilde Hutin retweetledi

[#phimscience] 🌾🦠New insight into the mechanisms by which #viruses adapt to #plant #immunity by Bonnamy et al 2022 #rice #NLRs @PHIM_research @LabDiade @ird_fr @INRAE_France @INRAE_DPT_SPE @EugenieHebrard @francois_sabot @HPidon
nph.onlinelibrary.wiley.com/doi/10.1111/np…
English

Today we had the chance to attend a presentation by Ms Bachabi from @AfricaRice. Management of the seed bank at AfricaRice, quarantine rules and seed health led to fruitful discussions between participants ! @PHIM_research @xanthomonads @Emersys_IRHS @ird_fr

English

Semaine de formation sur le diagnostic des bactéries pathogènes transmises par les semences au @isra_ceraas dans le cadre du GDRI #NSSN-X. @ird_fr @PHIM_research @xanthomonads @Emersys_IRHS
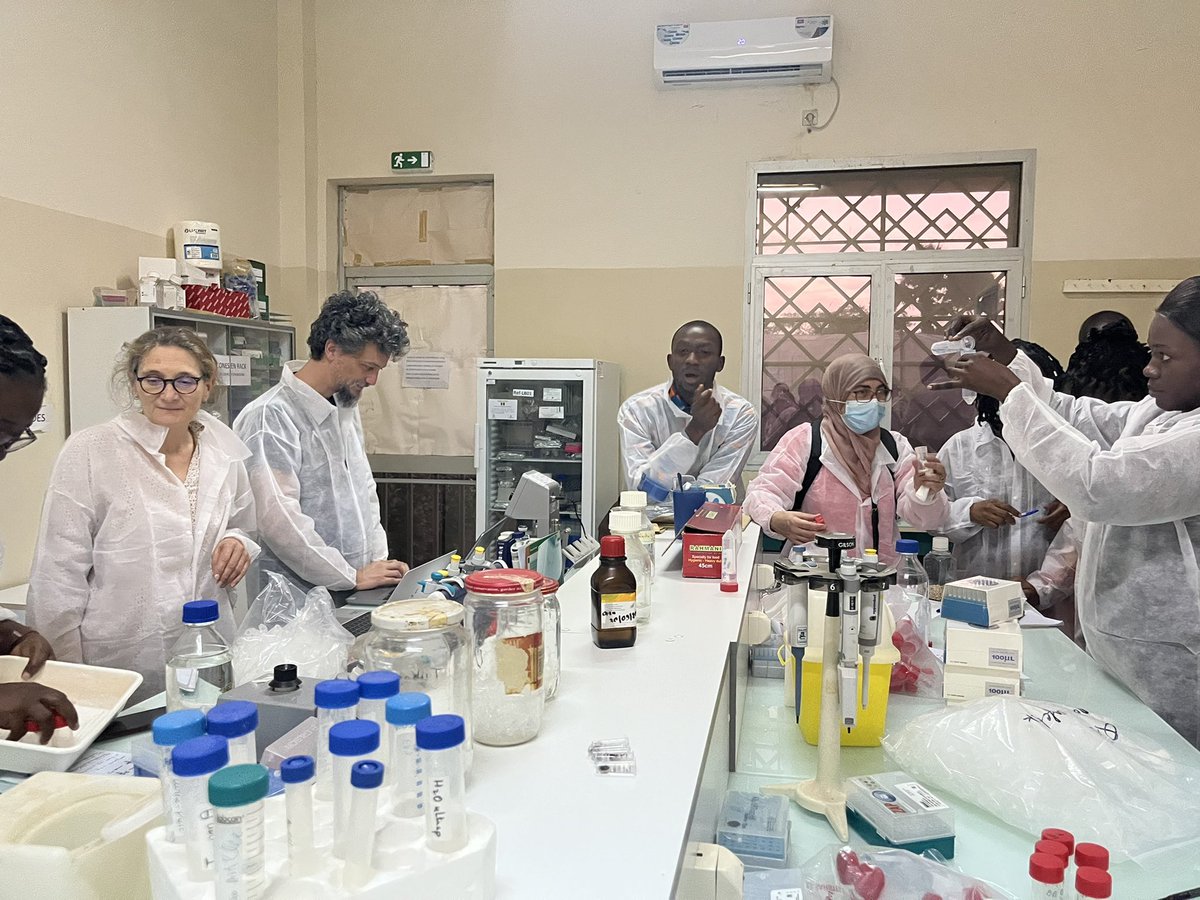
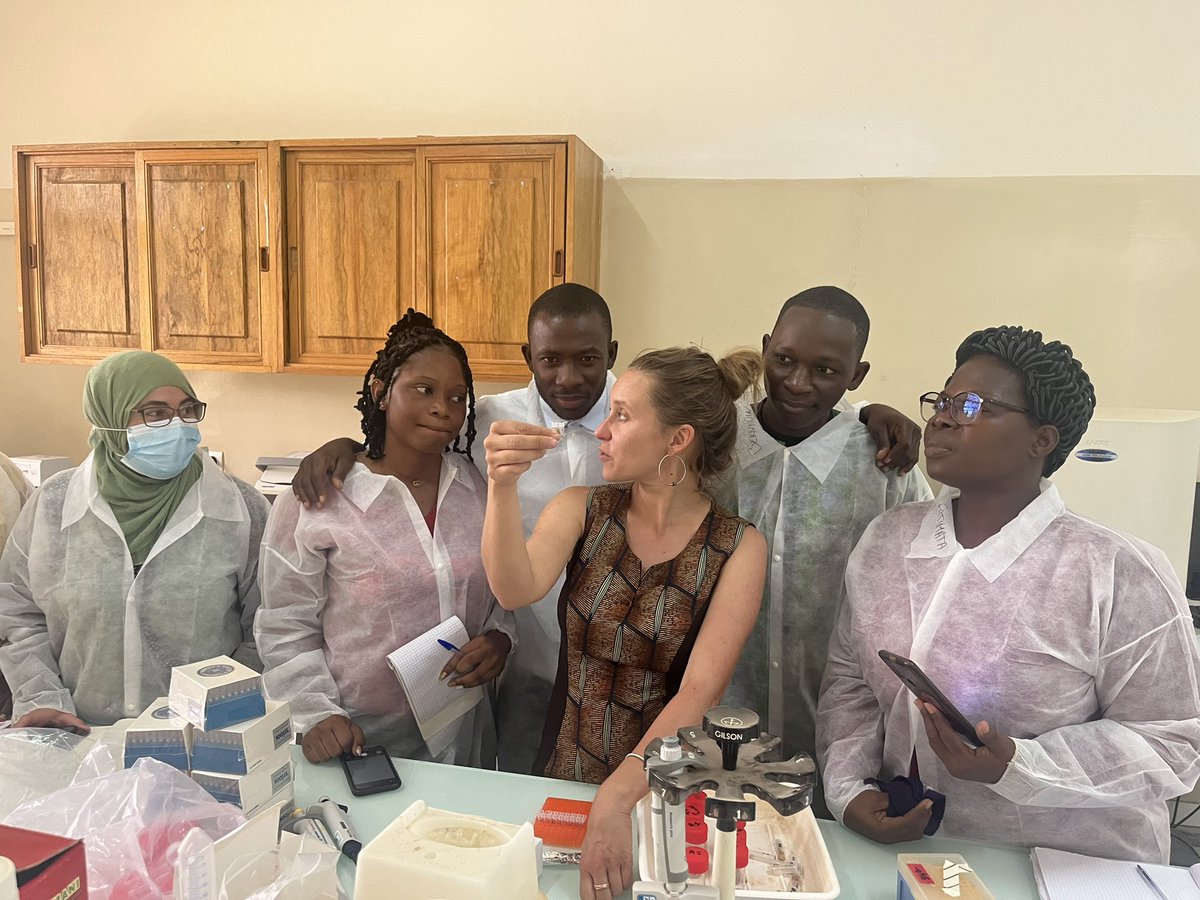



Français
Mathilde Hutin retweetledi

It’s my pleasure to announce the call for applications for the #NSSN-X Workshop on "Diagnosis of Seed-Borne Bacterial Pathogens" (in French) Participation in the course, including accommodation and meals, will be for free. Selected candidates may also profit from travel grants.

English
Mathilde Hutin retweetledi

TAL Effectors with Avirulence Activity in African Strains of Xanthomonas oryzae pv. oryzae | Rice | Full Text | @scoopit sco.lt/5qbVmi
English
Mathilde Hutin retweetledi

TAL effectors with avirulence activity in African strains of Xanthomonas oryzae pv. oryzae biorxiv.org/cgi/content/sh… #biorxiv_plants
Română
Mathilde Hutin retweetledi

Please RT. Phd funded for 3 years in my group on rice PGPR between France @ird_fr and Cambodia (ITC). Only for Cambodian citizenship students with a master (microbiology/ plant microbes interactions). More info drive.google.com/file/d/1ELERcm…
English
Mathilde Hutin retweetledi

Interesting preprint by Reshetnyak, @jmjacobs2 et al: #Xanthomonas infection in rice generates an atypical class of 20-22nt small RNAs that are often derived from protein kinase transcripts. Most of these xisRNAs do not seem to trigger cis-gene silencing. biorxiv.org/content/10.110…

English



