Maksim Kuznetsov retweetledi

I am excited to share that our BindGPT paper won the best poster award at @RealAAAI #AAAI2025! Congratulations to the team! Work led by @artemZholus!

Sarath Chandar@apsarathchandar
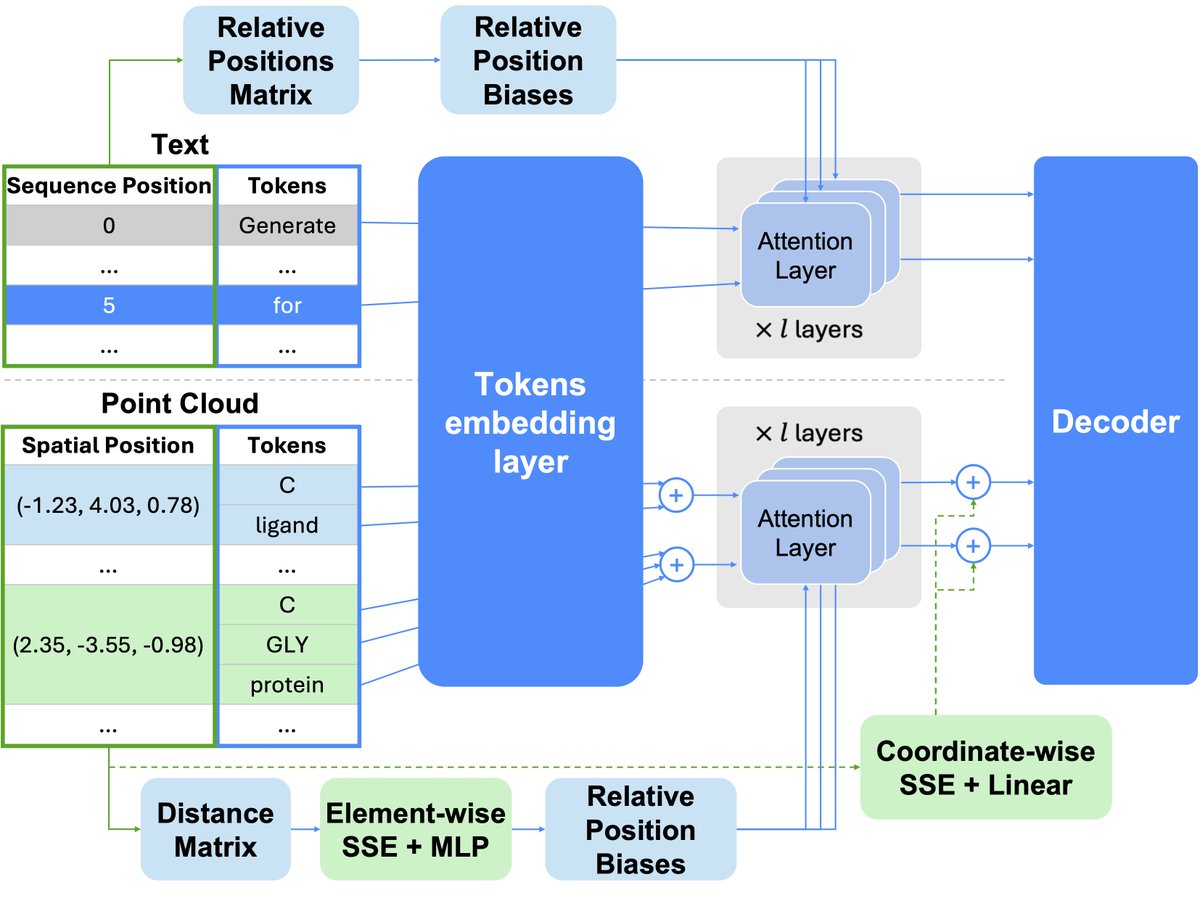
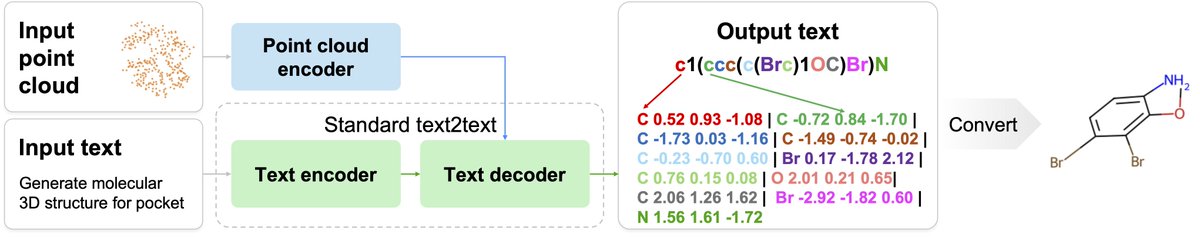
What's the foundational model for generative chemistry? Our work, BindGPT, is a good candidate, and it will be presented at #AAAI2025 today! We built a simple transformer language model that beats diffusion models by just generating 3D molecules as text! Led by @artemZholus 1/n
English