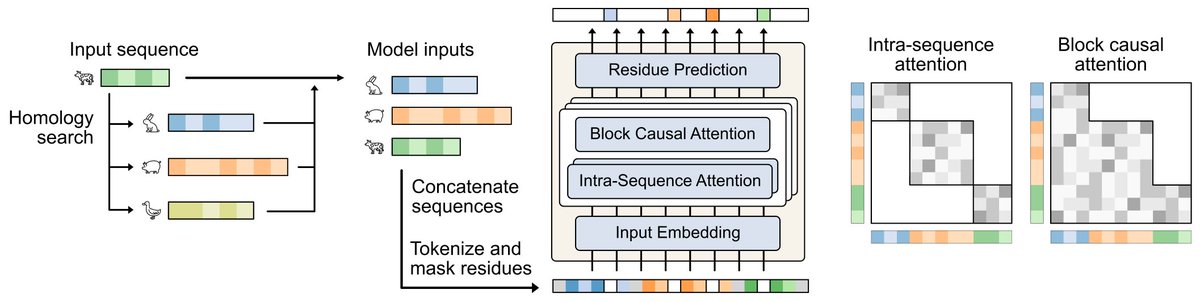
🚨ICML Paper Alert🚨 What if finding the right protein homologs wasn't a slow search, but a learned part of the model itself? We introduce 𝐏𝐫𝐨𝐭𝐫𝐢𝐞𝐯𝐞𝐫, an end-to-end framework that learns to retrieve the most useful homologs for self-supervised reconstruction! (1/12)