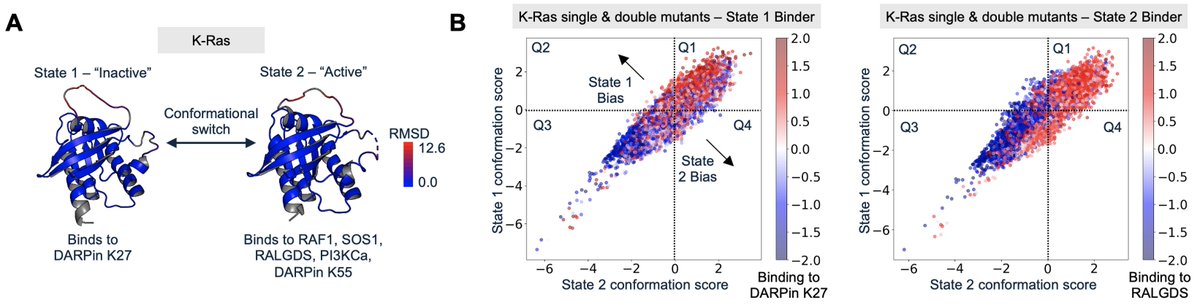
Can we design mutations that predictably bias proteins towards desired conformational states? Today in @ScienceMagazine, we introduce Conformational Biasing (CB), a simple and scalable computational method that uses contrastive scoring by inverse folding models to identify conformation-biasing mutations. science.org/doi/10.1126/sc…