
Markus J. Buehler
6.4K posts


Markus J. Buehler
@ProfBuehlerMIT
McAfee Professor of Engineering @MIT; Co-Founder & CTO at Unreasonable Labs; AI-Driven Scientific Discovery




AI x Life Science — the panel. Mod: Cheng Liu (Eureka Therapeutics) With: Ali Khademhosseini (3x Founder & CEO), Tim Kuruvilla (ex-Roche, SkyDeck), Deniz Kent (Prolific Machines), Hoifung Poon (Microsoft Research) Five operators at the frontier of AI and biology. One stage. Science x AI Summit. May 13. Register: sair.foundation/events/science…

Standard AI learns to imitate. We introduce a new framework that trains AI to make new discoveries in science + engineering. Learning-to-discover + open source LM led to: 🥇best new bound on Erdos min overlap problem 🥇fastest GPU kernels 🥇better single-cell denoising + more!






Yesterday at @BrownUniversity @ICERM's workshop on “Agentic Scientific Computing and Scientific Machine Learning” I spoke about “Adaptive Swarms Across Scales”, making the case for scientific AI as systems that can create representations, stress them, fracture them, and enlarge the category in which future representations live. The category here is a composable and breakable working universe of science: data, hypotheses, simulations, measurements, tools, failures, figures, papers, provenance, and the transformations that connect them. Discovery happens when those transformations become executable, inspectable, composable, and capable of changing the world model they operate within. Atomistic modeling gives one category - states, forces, trajectories, observables, boundary conditions, conservation laws. Neural surrogates learn fast morphisms inside or between such categories. But discovery is higher-order: it changes which objects and morphisms are available in the first place: what variables exist, what operations are allowed, what evidence counts, what scale is active, what invariant is being preserved, and what kind of explanation the system is even capable of forming. This is scientific method as adaptive architecture: compression, stress, fracture, recomposition. Fracture matters here because it makes the logic physical: a non-commuting diagram realized in matter. The imposed load, material hierarchy, defect field, and assumed continuum description no longer map cleanly into the observed outcome. The crack is the obstruction and it identifies where the old morphism failed and where a new representation must be introduced. The physical crack and the categorical obstruction are the same event viewed in different substrates. ScienceClaw × Infinite is a machine for constructing and transforming a category of scientific artifacts. Each artifact is typed. Each operation has lineage. Each failed branch remains in the category as reusable structure. The “paper” is no longer the terminal object of science; it is one projection of a larger compositional trace, and it can be generated at any time for consumption by a human or an AI. With that the unit of scientific labor is changing. For most of the twentieth century the unit was the result (a measurement, a theorem, a synthesized molecule). It is now becoming the algorithm that produces results, and after that, the substrate of discovery itself. The static PDF is the wrong terminal object for this regime, and the role of the scientist with it. We now design algorithms that build algorithms, and eventually substrates in which such algorithms compose themselves. At that point, the scientist is no longer outside the discovery system. The scientist becomes one of the representations the system can transform. In that sense, the systems will eventually do science to us, and that is the structural consequence of the principle they are built on.

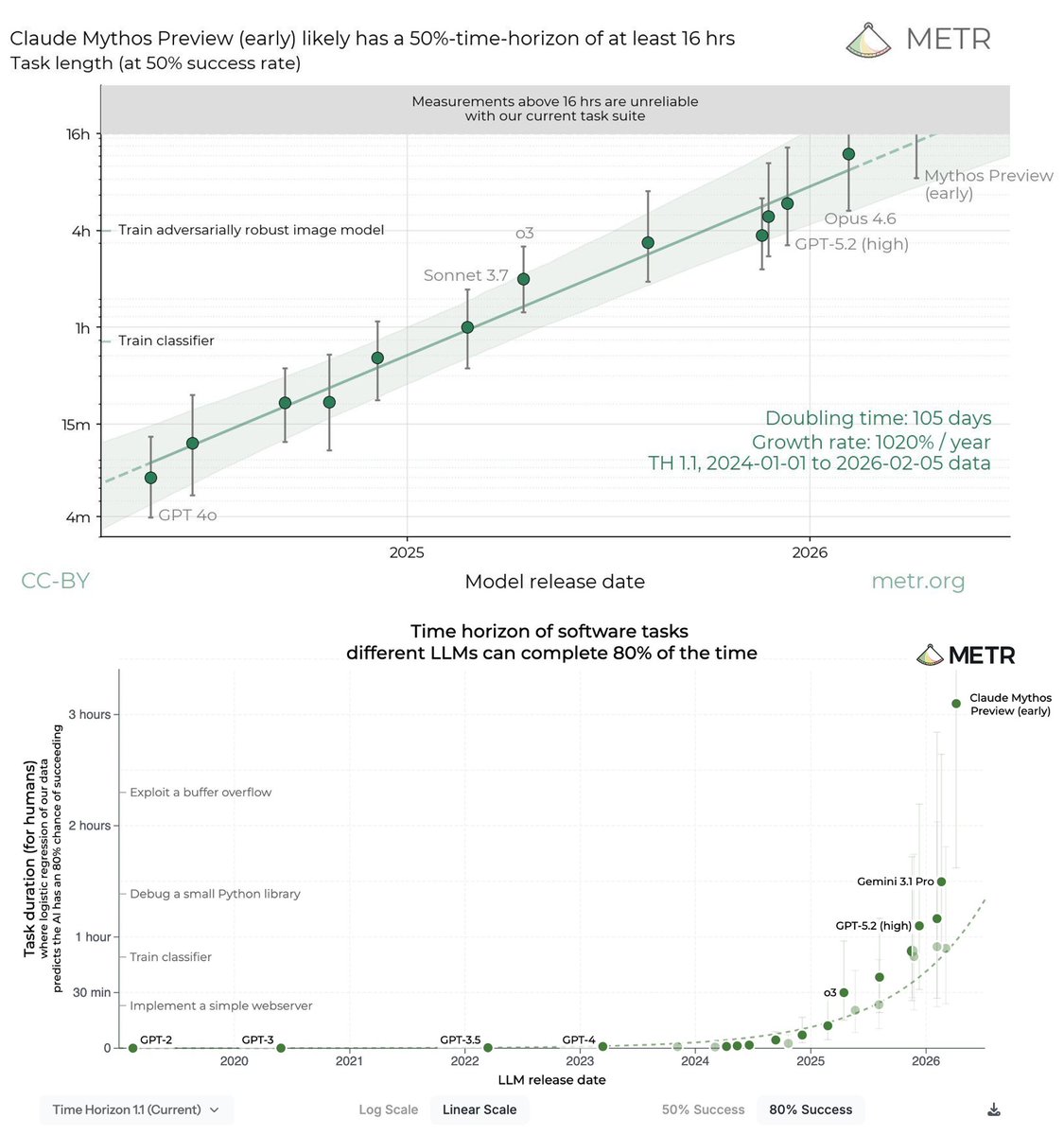

We evaluated an early version of Claude Mythos Preview for risk assessment during a limited window in March 2026. We estimated a 50%-time-horizon of at least 16hrs (95% CI 8.5hrs to 55hrs) on our task suite, at the upper end of what we can measure without new tasks.

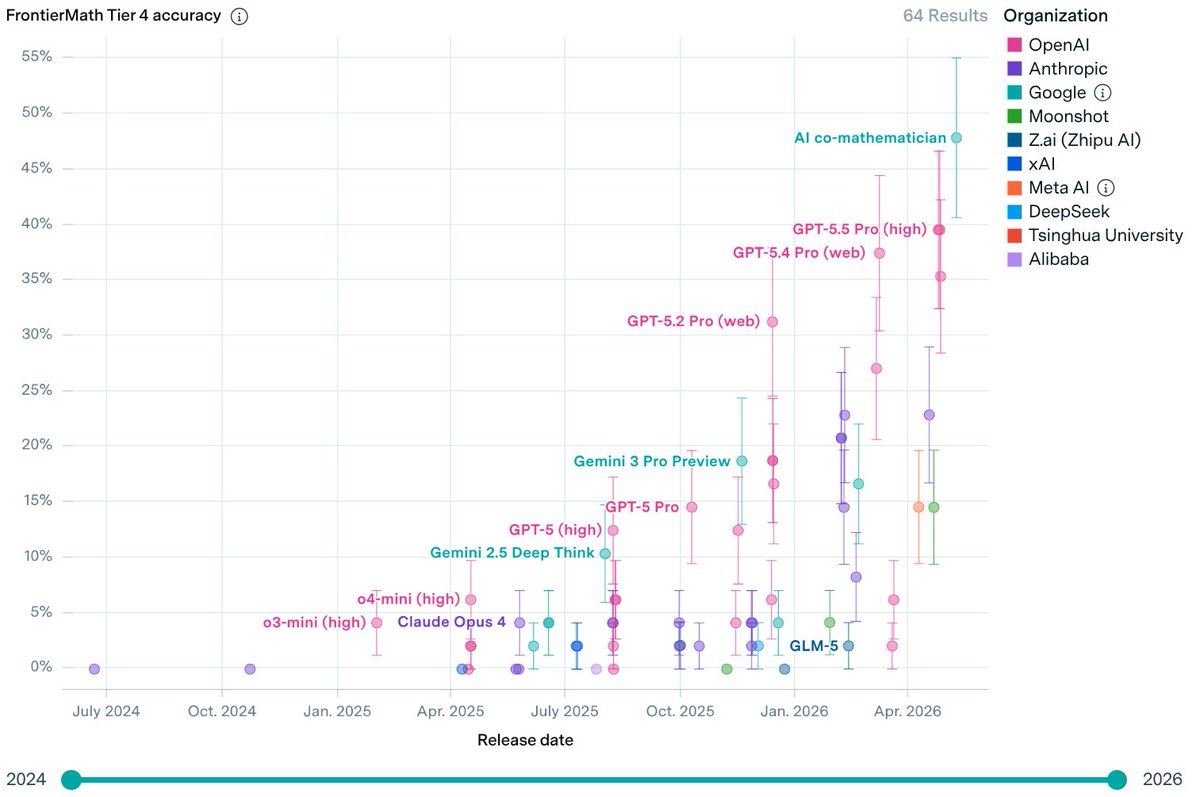

I've recently got in on the act of getting AI to solve open problems in mathematics. More precisely, I gave some questions asked by Melvyn Nathanson to ChatGPT 5.5 Pro, to which I have been given access, and it answered them. 🧵







I started Lambda in 2012 training ConvNets on an NVIDIA GPU workstation. Today, we’ve hit nearly a billion dollars in AI cloud revenue and provide compute to the most important companies in the world. Building a generational company is a lifelong quest and it’s all about the people who join you along the way. After 14 years as CEO, I’m returning to my roots to build Lambda as our CTO. I’m welcoming Michel Combes to join Lambda as our new CEO. Michel brings massive capital formation and allocation experience as the former CEO of some of the most storied infrastructure companies in the history of the world: SoftBank International and Sprint. It’s been an honor serving the AI community as Lambda’s founding CEO and I’m looking forward to continuing to serve you from the CTO seat! I’m excited to work alongside Michel and everybody at Lambda to build an iconic 100-year company.


