Salzman Lab
58 posts


Salzman Lab
@SalzmanLab
Official Twitter of the Salzman research group at Stanford Biomedical Data Science
Stanford, CA Katılım Haziran 2017
22 Takip Edilen302 Takipçiler

presentation by @TBaharav today at @RECOMBconf on the statistics behind SPLASH, a method that can be used as a robust alternative to one of the most common statistical tests
English

backlogged on posting our new work in genomics! using RNA splicing as a special case of expansive, ultra-efficient and fully statistical analysis of RNA or DNA sequencing data. cell.com/cell/fulltext/…
English
Salzman Lab retweetledi

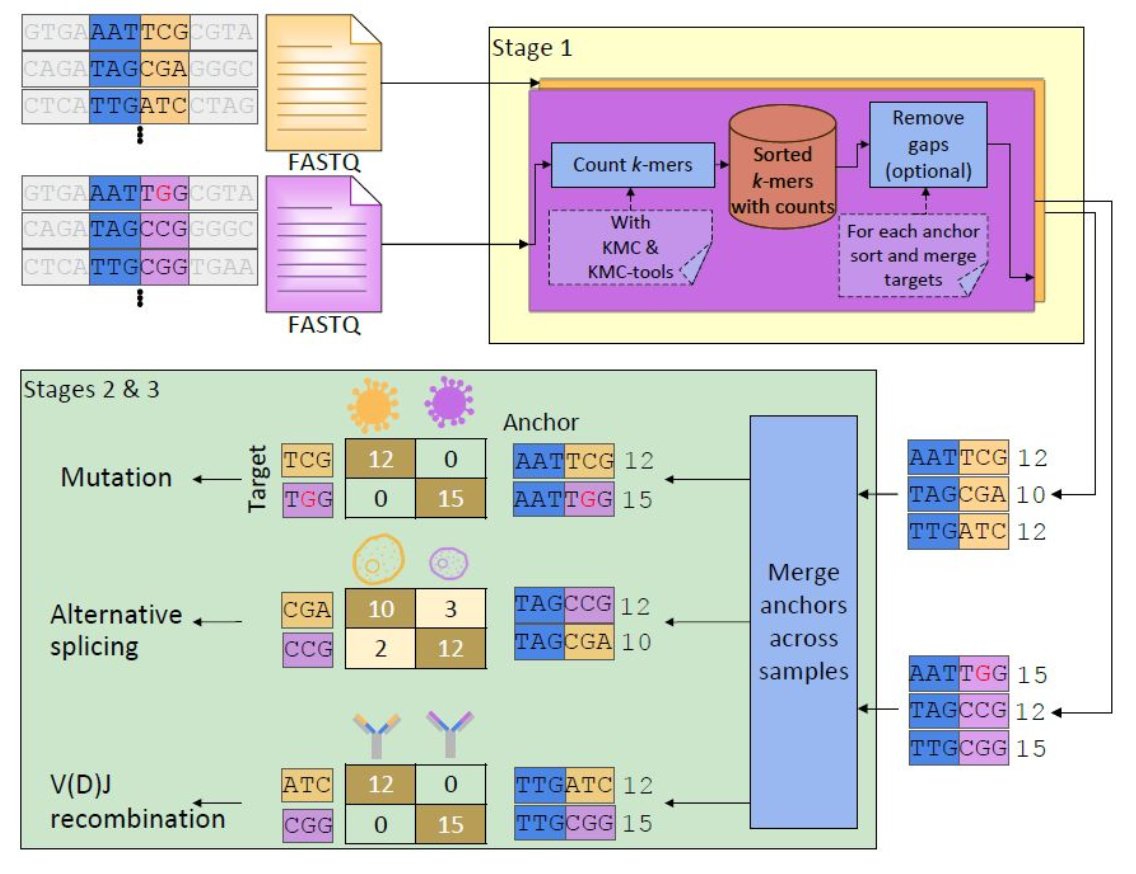
3. Development of NOMAD2, which enables reference-free biological discovery at unmatched scale and speed. In collaboration with @sdeorowicz and @marekkoki, led by @marekkoki and @roozbehdn, this package is publicly available on github: github.com/refresh-bio/R-…
English

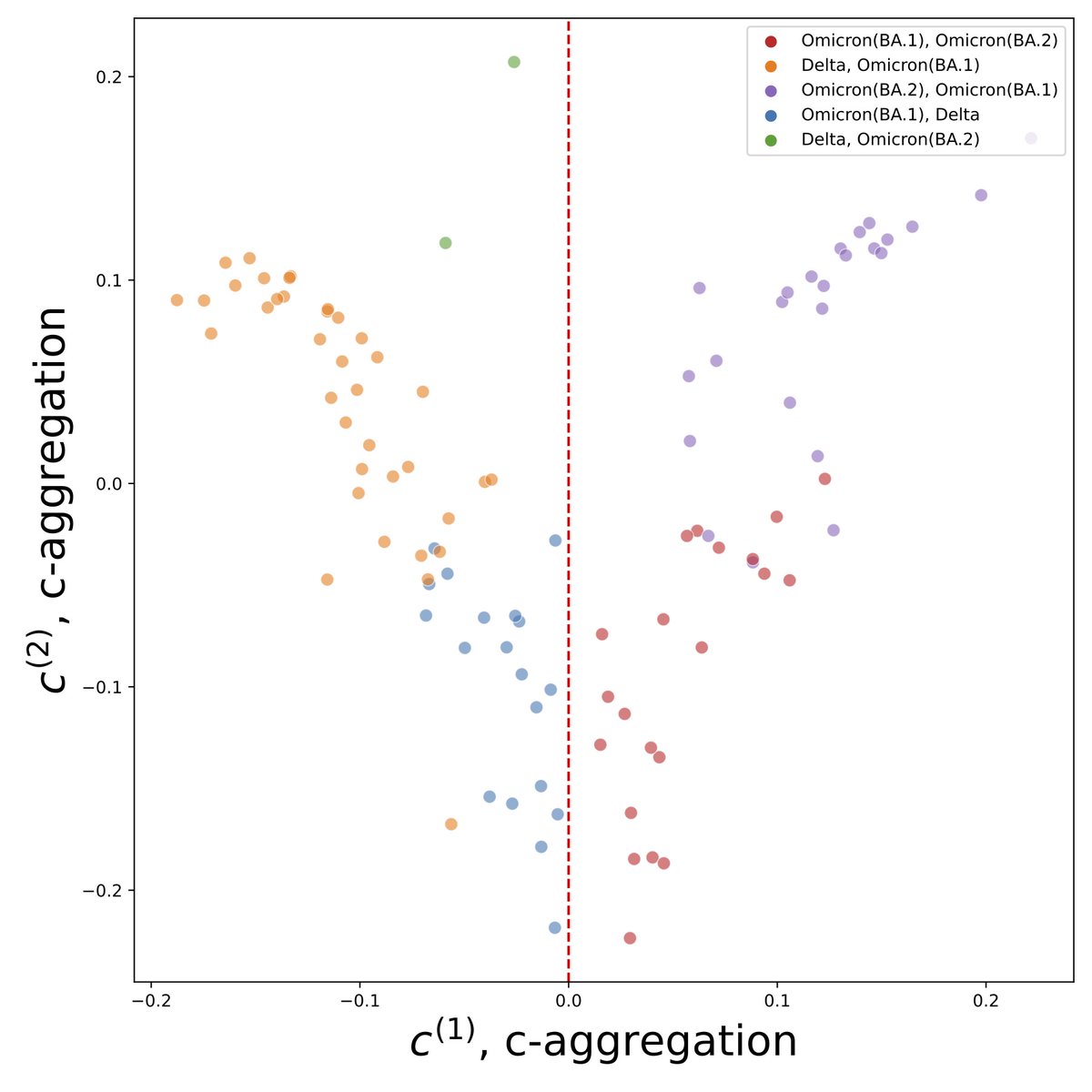
2. Design of a novel statistical test on contingency tables, OASIS, which provides interpretable and finite-sample valid rejection of the null, enabling reference-genome and metadata free strain detection in biological applications. Led by @TBaharav: biorxiv.org/content/10.110…
English

Thrilled to announce 3 new papers from our lab, up on bioRxiv this past week.
biorxiv.org/content/10.110…
biorxiv.org/content/10.110…
biorxiv.org/content/10.110…
English
Salzman Lab retweetledi

I can confirm that the NOMAD analysis method and the accompanying software is a game-changer for sequence analysis and discovery. Check this paper out! biorxiv.org/content/10.110…
English
Salzman Lab retweetledi

NOMAD2 is an unsupervised, reference-free, ultra-fast next generation of NOMAD, revealing new insights into cancer transcriptomes. Joint work with @salzmanlab. Many thanks to all for great collab!
biorxiv.org/content/10.110…
English

Excited to share that our paper on reference-free transcriptomic analysis is live. biorxiv.org/content/10.110…
English
Salzman Lab retweetledi

Had a blast at #biodata22! Was great to present and get feedback on NOMAD, our recent work for reference-free inference biorxiv.org/content/10.110….

English
Salzman Lab retweetledi

"NOMAD is a single reference-free ‘omic algorithm for biological discovery across the tree of life"
Julia Salzman @SalzmanLab en #FrontiersInGenomics HOY martes 15 nov 5pm #EnVivo #EnLínea lcg.unam.mx/frontiers/live @ccg_unam @ibt_unam @unammorelos

English

Very excited that ReadZS, a scRNA-seq pipeline for detection of RNA processing, is out in @GenomeBiology! This work was led by @ea_meyer, K Chaung, and @roozbehdn.
Genome Biology@GenomeBiology
ReadZS, from @ea_meyer, Chaung, @roozbehdn & Salzman, for identifying RNA processing (splicing, alternative polyadenylation etc) in RNA-seq data, without the need for pre-existing annotations. It works on both 3'-enriched and non-enriched data genomebiology.biomedcentral.com/articles/10.11…
English

Excited to share our new preprint NOMAD! We present a unifying statistical formulation for many fundamental problems in genome science and develop an efficient reference-free algorithm (NOMAD) to solve it. Work by Kaitlin Chaung, @TBaharav, and @RZheludev. biorxiv.org/content/10.110…
English

Excited to announce that a project spearheaded between our lab and David Tse's has been selected for a @czbiohub Data Insights Award! Follow the link for more information.
chanzuckerberg.com/science/progra…
English

Excited to share DIVE is live on bioarxiv! DIVE is a novel reference-free approach for de novo discovery of diversity-generating and mobile genetic elements implemented as a publicly available python package (work led by @jordiatjhu).
bioRxiv@biorxivpreprint
DIVE: a reference-free statistical approach to diversity-generating and mobile genetic element discovery biorxiv.org/cgi/content/sh… #bioRxiv
English