Sabitlenmiş Tweet

New vcfdist paper on bioRxiv!
Key Takeaways:
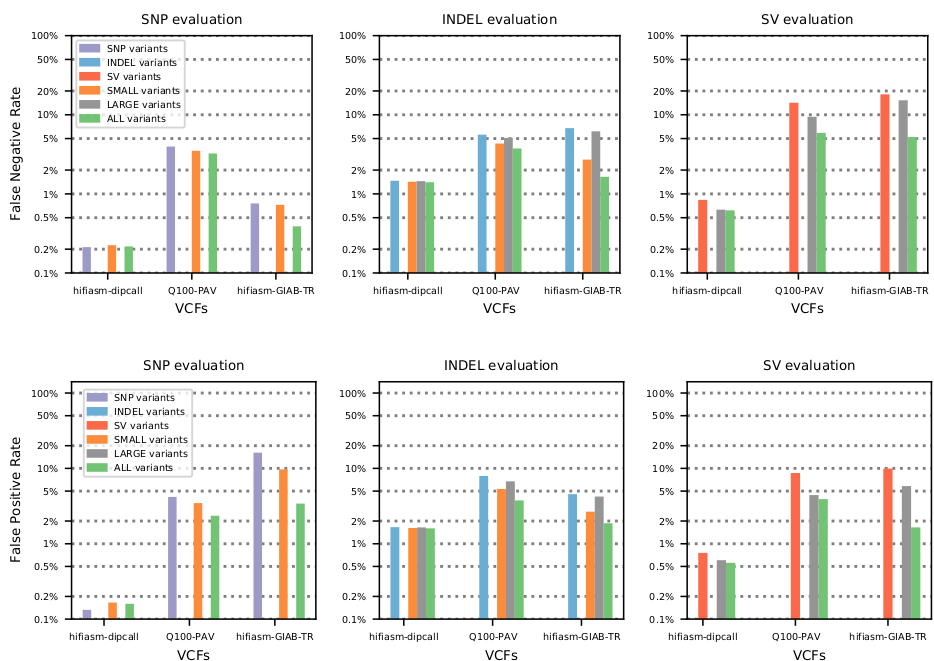
1) Jointly evaluating small and structural variants decreases measured FN+FP by 20-50%
2) 43-92% of phasing flip "errors" are FP due to variant representations
More below
Code: github.com/TimD1/vcfdist
Paper: biorxiv.org/content/10.110…
English