Our latest from @TheCrick, now on @biorxivpreprint: biorxiv.org/content/10.110… Michael Herger (@HergerMichael) and Christina Kajba jointly led development of a new prime editing platform for testing large numbers of human genetic variants.
Greg Findlay
281 posts


@TheGenomeLab
Group Leader of the Genome Function Laboratory @TheCrick From 2025, find me over on the other side: https://t.co/ajOqpCvaUY (he/him)

Our latest from @TheCrick, now on @biorxivpreprint: biorxiv.org/content/10.110… Michael Herger (@HergerMichael) and Christina Kajba jointly led development of a new prime editing platform for testing large numbers of human genetic variants.







(1/4) I’m delighted to announce that our research on RNU4-2 is now out on @Nature . This is an exciting finding that will bring many diagnoses worldwide. We have updated some new results since the preprint: nature.com/articles/s4158…



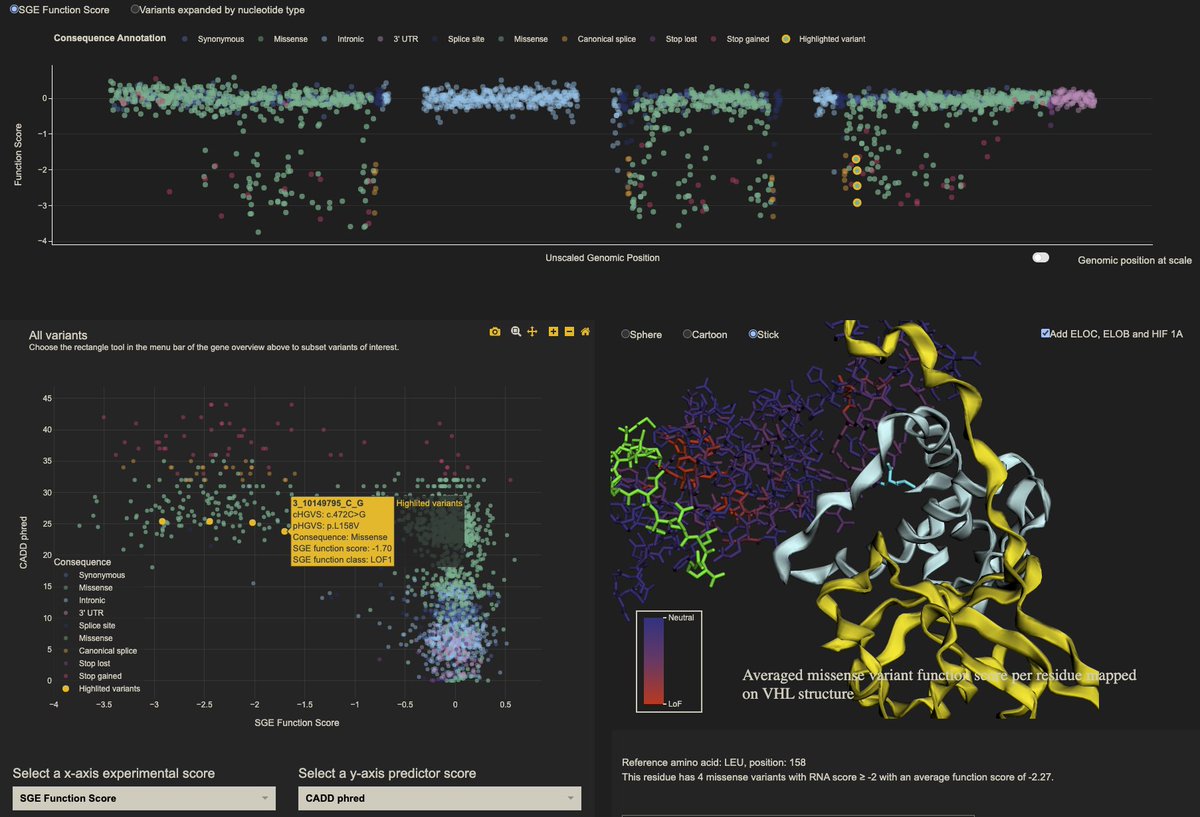
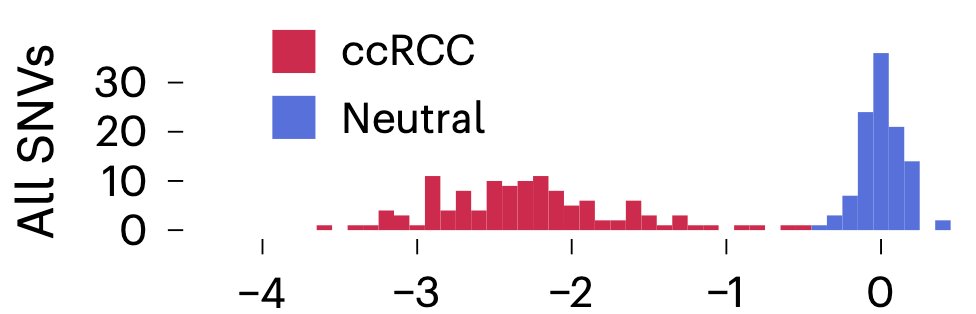
The VHL data can also be searched, visualised, and explored using this awesome SGE visualisation platform made by @ChloeTerwagne: vhl-board.onrender.com Chloé’s code is all on GitHub if you’re interested in viewing and sharing your own data like this. (3/n)