WashUDevBio
209 posts





The June issue is live nature.com/nbt/volumes/42… On our cover, Roohani et al. present GEARS, a computational method that integrates deep learning with a knowledge graph of gene–gene relationships to simulate the effects of genetic perturbations go.nature.com/3E37kEo

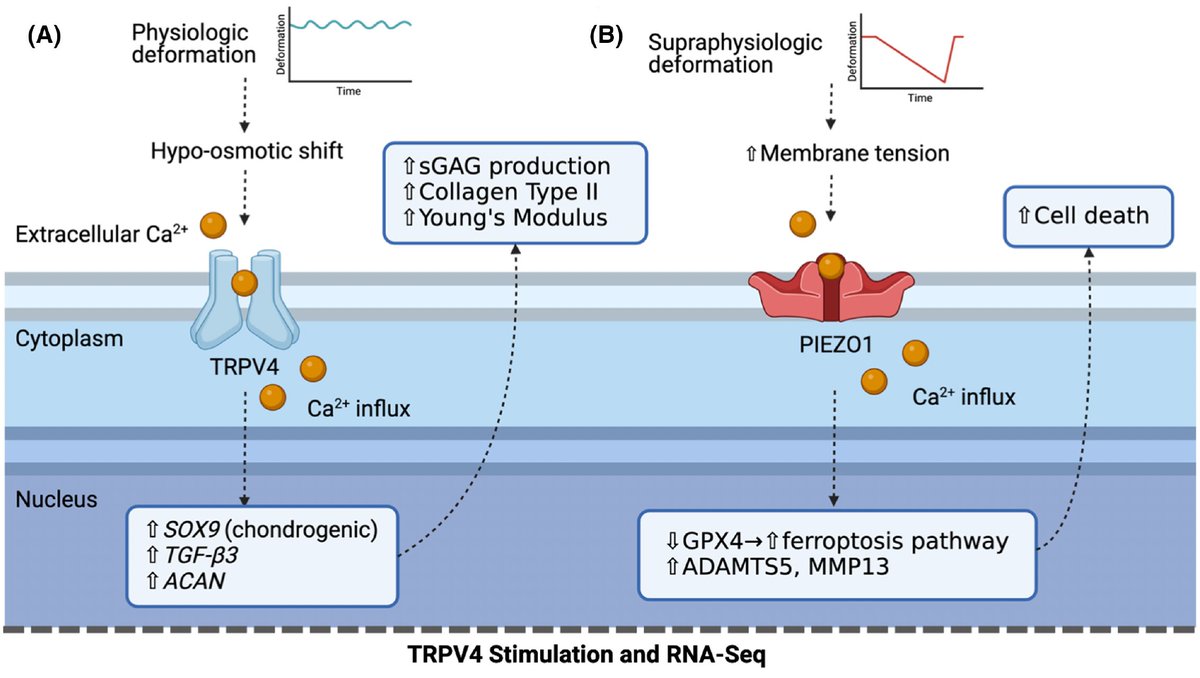

Research from WashUMedicine in collaboration with @UFexplore reveals how brain inflammation triggers extreme muscle weakness across several diseases, including viral infection, bacterial infection and Alzheimer’s disease. medicine.wustl.edu/news/brain-inf…

In this Spotlight, @HsiuChuan_Lin (@ETH_BSSE) @AlyMakhlouf93 (@MRC_LMB) @CamilaVazquezE (@lundu) Dorota Zawada @TU_Muenchen & @simoesfili discuss the main points emerged from @Co_Biologists Workshop ‘Novel Technologies for Programming Human Cell Fate’: journals.biologists.com/dev/article/15…


Thrilled to share the final publication of CellTag-multi, a strategy in which CellTag lineage barcodes can be captured in both scRNA-seq and scATAC-seq assays allowing for independent tracking of clonal transcriptional and epigenomic state, with @kjkjindal twitter.com/NatureBiotech/…

Join us Thursday, October 5th at 4:30pm in the Connor Auditorium for the 42nd annual Oliver H. Lowery Lecture with speaker Didier Stainier.



Single-cell lineage capture across genomic modalities with CellTag-multi reveals fate-specific gene regulatory changes go.nature.com/3rkazov












