Sabitlenmiş Tweet
William Pearman
193 posts

William Pearman
@WilliamPearman
Research fellow at Auckland Uni. Same handle at the blue place. Evolutionary ecology, microbial ecology. Anglican. He/him
Tāmaki Makaurau, New Zealand Katılım Mayıs 2017
659 Takip Edilen283 Takipçiler
William Pearman retweetledi

Joint consideration of selection and microbial generation count provides unique insights into evolutionary and ecological dynamics ... biorxiv.org/cgi/content/sh… #biorxiv_micrbio
English

Glad to have this paper out now! If you've ever tried and less-than-successfully sequenced brown algae - polyphenols might be why!
onlinelibrary.wiley.com/doi/full/10.11…
English

Looking for some @nanopore help - I have some ~1600 bp amplicons sequenced with the rapid kit (plasmidseq type data) and want to cluster them into ASVs/OTUs - any idea how? Mixed of read lengths due to data type, and only have 2 samples.
English

@JLGONZHER @environmicrobio @kirk3gaard @nanopore Stay tuned! I've got some ideas on this that I'm hoping to try out!
English

@WilliamPearman @environmicrobio @kirk3gaard @nanopore Thank you for sharing. Now we just a need a good way to clean the samples
English

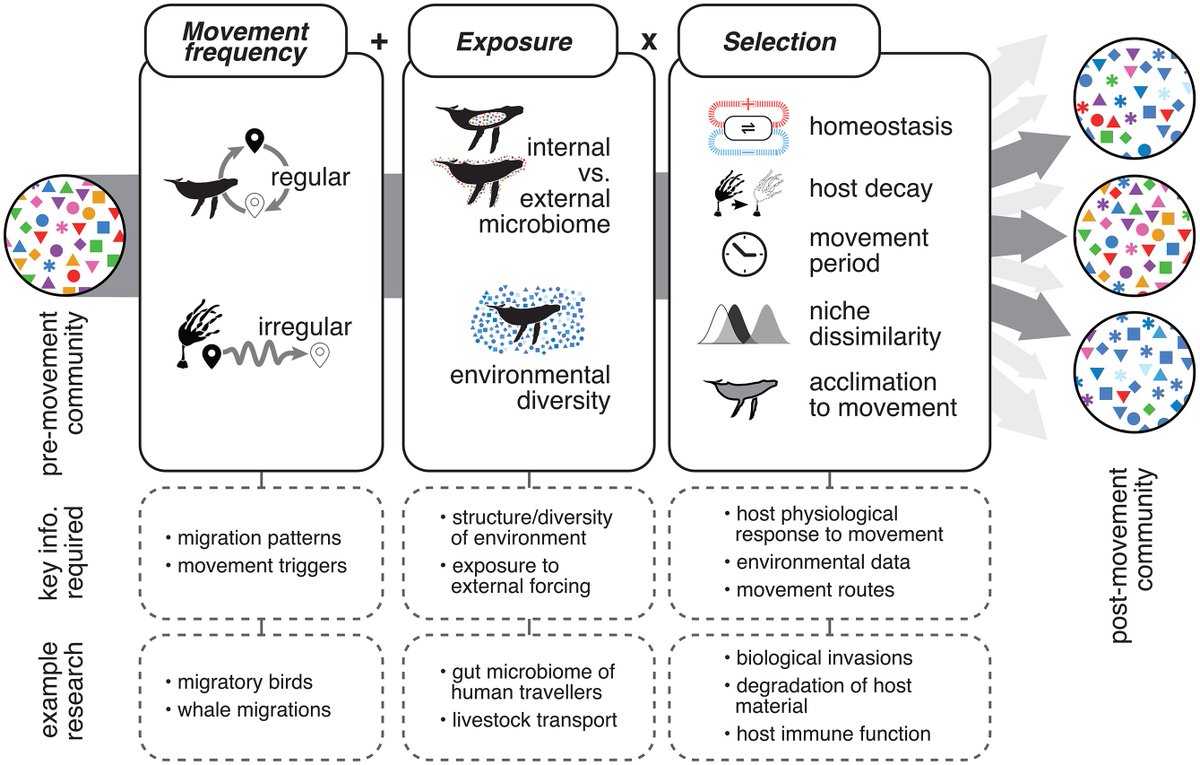
2/2 - so we wrote up a piece on this, developing models and hypotheses about what we expected would be the case, differences to migratory organisms etc. Hugely grateful to my fantastic co-authors, esp @Grant_Duffy who made the wonderful figure above!
English

Glad to have this perspective piece out now! We found that when we were trying to develop hypotheses around how microbiomes might change in response to host dispersal - there wasn't much information for non-migratory organisms (e.g., kelp) 1/2
academic.oup.com/femsec/article…
English

Glad to have this neat little pre-print out with a bunch of great folks! @NickGreen_chem @Evie_Jensen
biorxiv.org/content/10.110…
After struggling so much to get good sequenceable DNA from seaweeds - we did some digging and found some cool results!
English

Note the 260/230 and 260/280 ratios for these samples were 'good' - and yet seemingly have a bunch of other stuff in there! This was my first time working on this sort of data, so a huge thanks to @NickGreen_chem for being a great guide into the world of chemistry!
English

@JLGONZHER @environmicrobio @kirk3gaard @nanopore Just following on from this, here's the pre-print.
biorxiv.org/content/10.110…
English

@JLGONZHER @environmicrobio @kirk3gaard @nanopore Any idea how the tissue was stored prior to extraction? I think one that's becoming apparent is that for plants/algae - minimizing storage time is essential.
English

@WilliamPearman @environmicrobio @kirk3gaard @nanopore I had the same happened when trying to sequencing a grass genome; pores would die on ~ 6hours
English

@JLGONZHER @environmicrobio @kirk3gaard @nanopore I'm currently working on some ideas in that vein, but at this point we've used LC-MS to show clear polyphenol contamination that wasn't seen with abs ratios. I'll link the pre-print in this thread once it's up in a few days.
English

@WilliamPearman @environmicrobio @kirk3gaard @nanopore Any suggestions on additional quality/purity evaluation beside the abs ratios?
English

@JLGONZHER @environmicrobio @kirk3gaard @nanopore 2/2 Even 1 polyphenol adducted base in 10,000 could be problematic - but not detected with abs ratios. I've got a pre-print coming out this week hopefully describing the problem.
English

@JLGONZHER @environmicrobio @kirk3gaard @nanopore Even abs ratios can be misleading, when we've worked on macroalgal samples - we very rarely get good sequencing results despite 'perfect' input DNA.
Largely we think because polyphenols can bind to the DNA at low levels - and lead to pore damage. 1/2
English

Happy to have this paper out this week! With @Clare_eDNA @CeridwenFraser Fine-scale phylogenetic diversity gradients support the Antarctic geothermal refugia hypothesis | Antarctic Science | Cambridge Core - bit.ly/4bvkHMf
English
William Pearman retweetledi

Excited to share our new paper @EcographyJourna, where we use population genomics to investigate the impact of the 2016 Kaikōura earthquake on Durvillaea #kelp/rimurapa!
First genomic snapshots of recolonising lineages following a devastating earthquake: doi.org/10.1111/ecog.0…

English