Ya (Allen) Cui
55 posts

Ya (Allen) Cui
@YaCui7
Assistant Professor of Bioinformatics, School of Medicine, UC Irvine Lab website: https://t.co/9I2KFXsflY
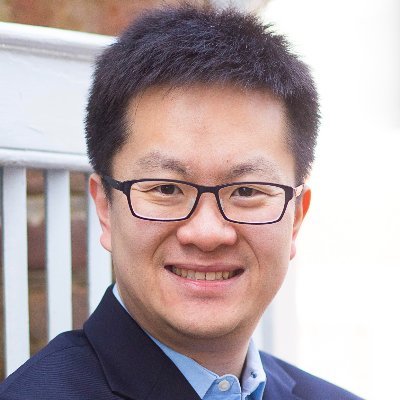


Congratulations to Dr. Chongzhi Zang, a recipient of the 2024 Dean's Award for Excellence in Research. Thank you for all the great work you do!



Congratulations to the 2024 Class of Pinn Scholars: Jennifer Payne, MD, Jie Sun, PhD, and Chongzhi Zang, PhD. Pinn Scholars are mid-career faculty chosen for their scientific expertise and research productivity. @zangcz #UVA bit.ly/4d1kd1t
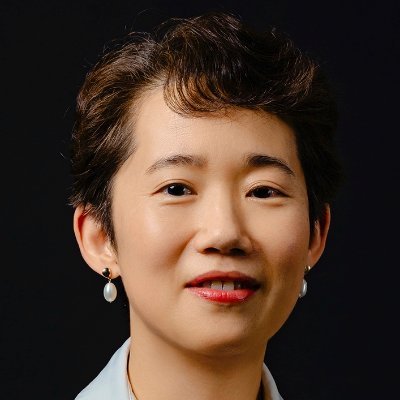



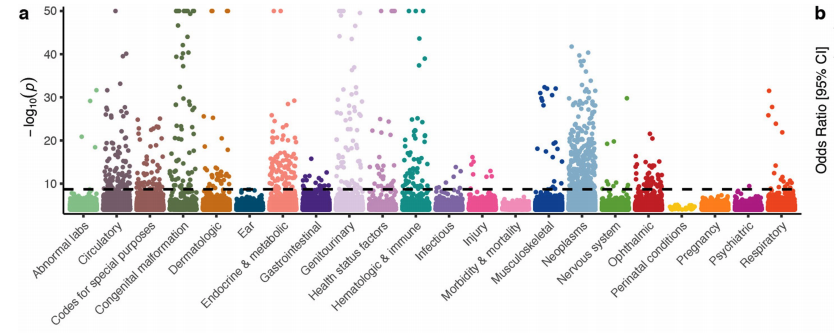
Now online! A genome-wide spectrum of tandem repeat expansions in 338,963 humans dlvr.it/T576KX


Our new paper @NatureComms performed the first Alternative polyadenylation transcriptome-wide association study for 11 brain disorders with 17,300 RNA-seq samples. We identified 354 novel APA-linked risk genes including ATXN3 for ALS. nature.com/articles/s4146…
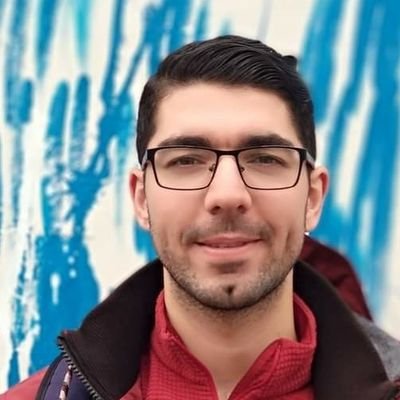
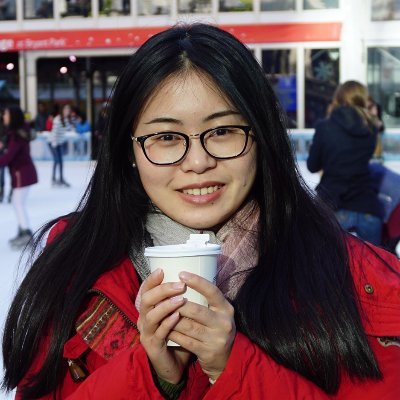





MAAPER: model-based analysis of alternative polyadenylation using 3′ end-linked reads, from @vivianstats, @BinTian_lab &co. It can be applied to reads close to, but not spanning, the polyadenylation site, and works with both bulk and single-cell data genomebiology.biomedcentral.com/articles/10.11…