Arman Seyed-Ahmadi
27 posts

Arman Seyed-Ahmadi
@arman1sa
AI/ML for CompBio @UHNAIHUB Previously @Genentech | PhD @UBC

Great to see 2 fabulous papers using model interpretation methods developed in our lab for deep learning models of regulatory DNA being to reveal causal mechanistic role of sequence syntax mediated TF binding, methylation & histone variants in stem cell memory of inflammation 1/

Proteins can now talk. Introducing BioReason-Pro, the first reasoning model for protein function. A thread🧵













Today we’re announcing X-Cell — Xaira’s first step toward a virtual cell. 🧬 A foundation model that predicts how gene expression changes under causal perturbations — across cell types, conditions, and even unseen biology. This is not trained on observational atlases. It is trained on interventions. 🧵👇

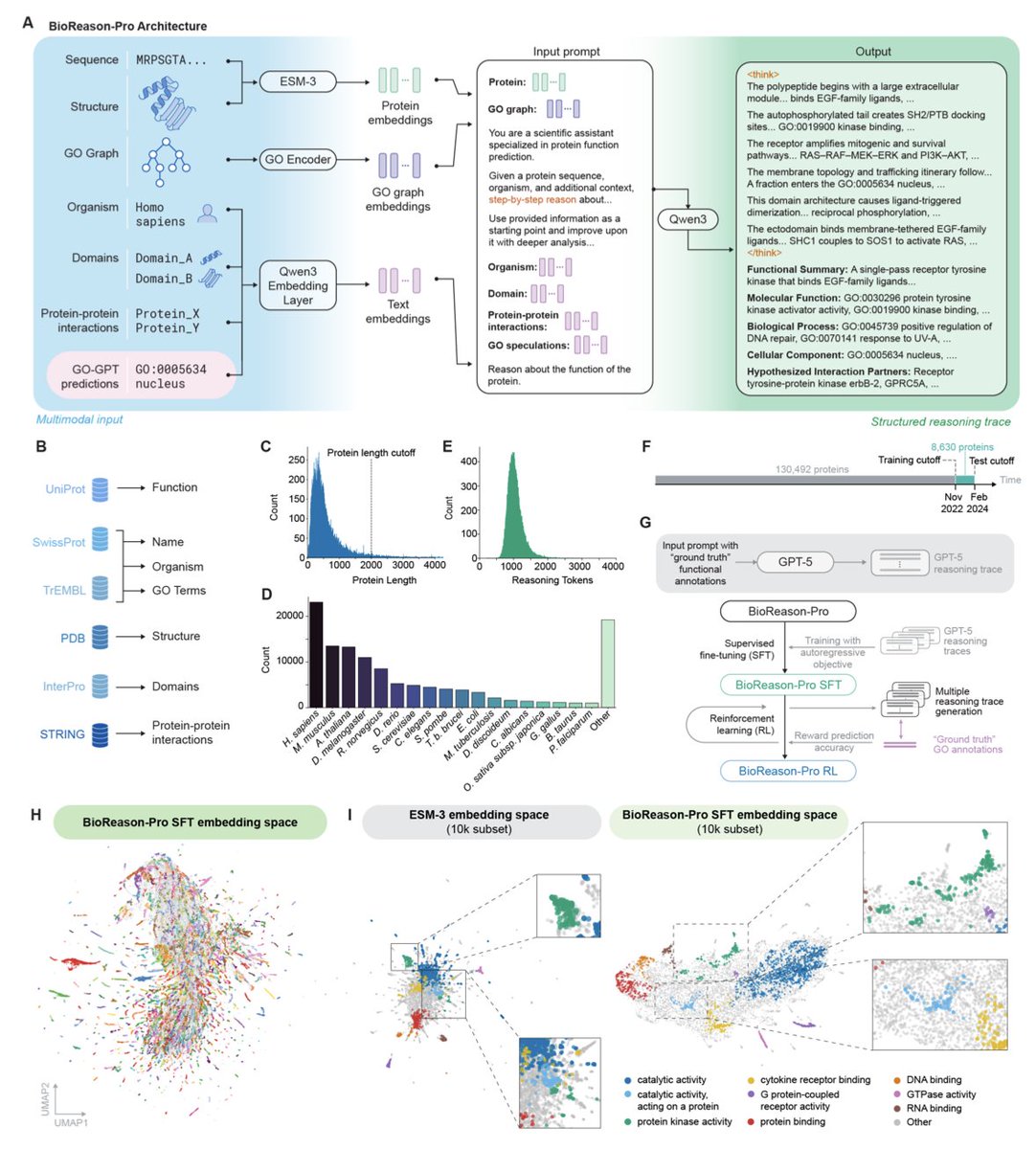
2026 may be the year AI starts to truly reason about biology. AlphaFold helped close the sequence → structure gap. The next frontier is sequence → functions. Today, together with @genophoria and the team at @arcinstitute , we’re releasing BioReason-Pro — the first multimodal reasoning model for protein function prediction.


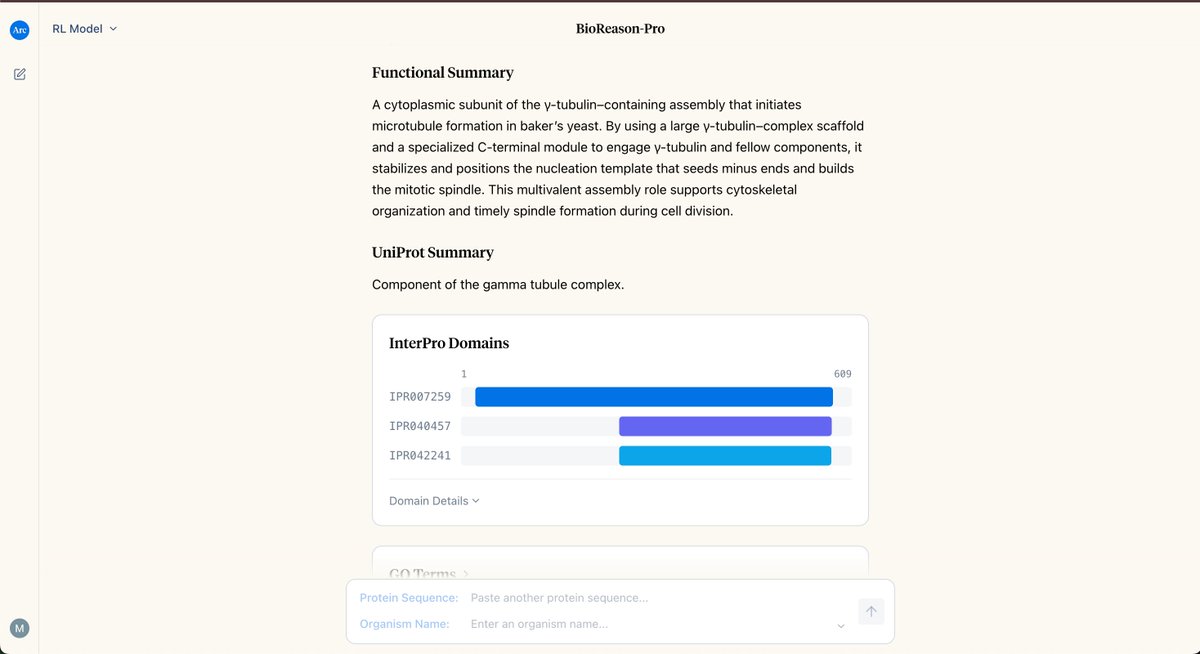
What if AI could explain why a protein is a kinase, not just tell you it is? We built just that. BioReason-Pro is a multimodal LLM that reasons about protein function — walking through domains, interactions, and biological context to make predictions you can actually evaluate.



Proteins can now talk. Introducing BioReason-Pro, the first reasoning model for protein function. A thread🧵


today we launched bioreason-pro, try using it: app.bioreason.net


