
Benjamin Furtwängler
43 posts



Only one more month until this year's iSCMS conference, hosted in Copenhagen, Denmark. Registration (and abstract submission in fact) is still open (singlecellms.org), so reserve your seat(s) now! :) Preliminary program now live too, with a stellar speaker lineup. #scMS
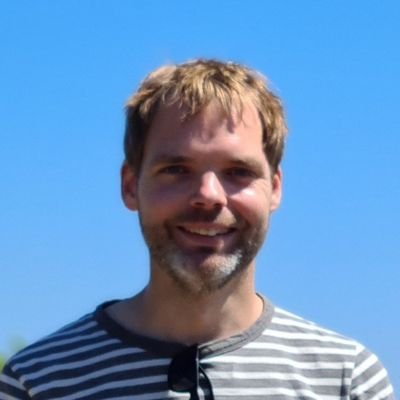




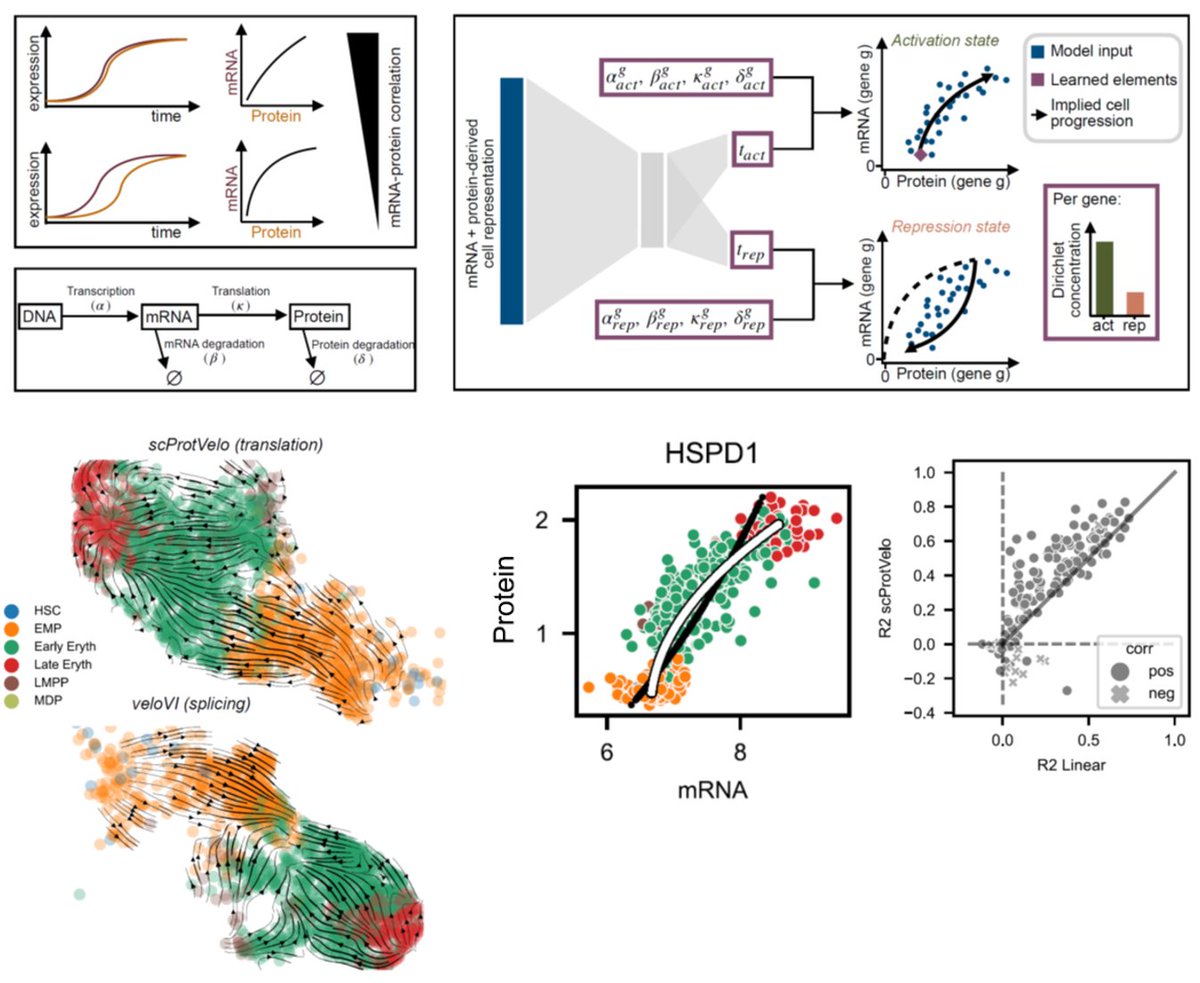
We’re excited to present this integrative analysis of single-cell proteomics and transcriptomics of the human HSPC hierarchy. Amazing teamwork with @NilUresin, @_sabrinarichter, @fabian_theis, @erwinschoof, @Bo_Porse biorxiv.org/content/10.110… 🧵




Have you ever wondered if you can tailor your DIA method to facilitate deeper proteome coverage from ultra-low-input samples? Well, look no further, as I am overjoyed to share our latest pre-print where we do precisely that. biorxiv.org/content/10.110…


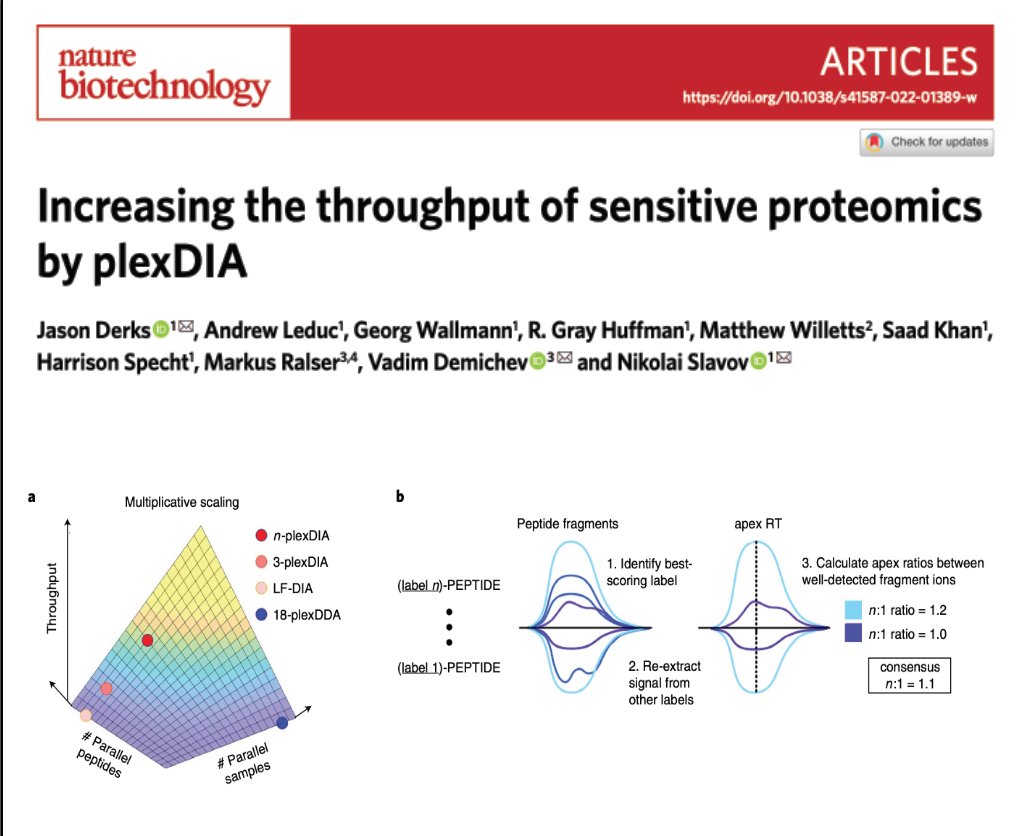
Proteomics should be sensitive, accurate, fast, robust & affordable. 👉 Welcome plexDIA! Proteins / sample: ~ 8,000 Sample size: 500 ng Samples / day: 48 (should be 72 with Evosep) MS system: 1st generation QE classic biorxiv.org/content/10.110…

