Cagla Eroglu
1.3K posts
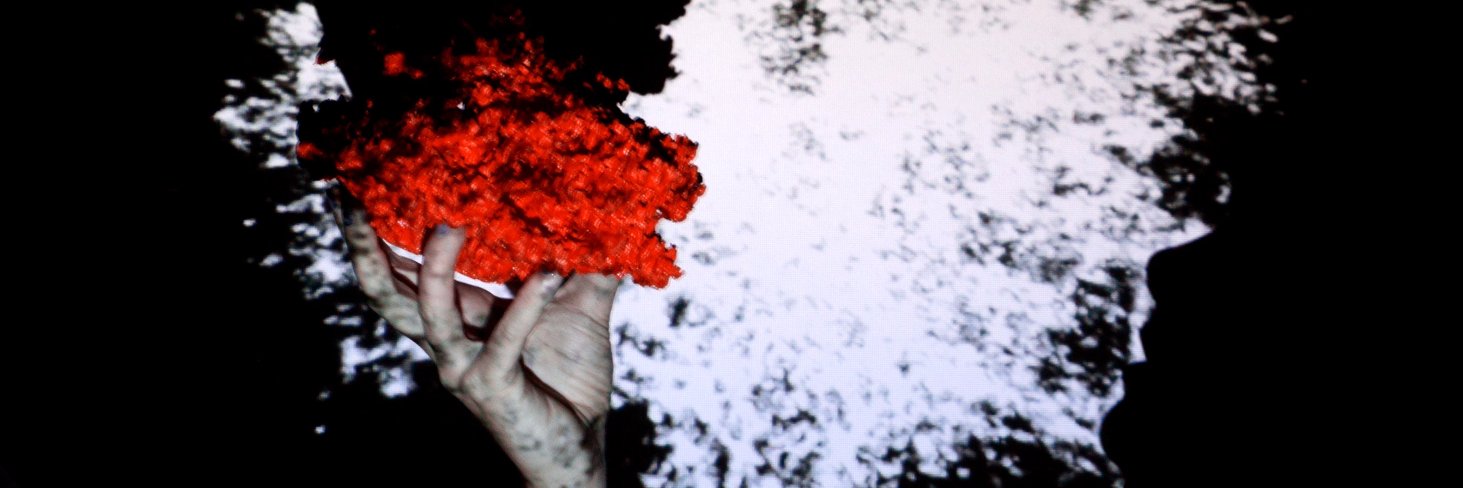







I am very excited about our new paper just published in PNAS. Dense Associative Memory is a versatile family of models that allows to store large amounts of information, have strong error-correcting capabilities, and other desirable properties. But is it possible to build it in biological “hardware”? This is a tricky question since DenseAMs have many-neuron couplings (more than 2 neuron couplings). This, at face value, may seem contradictory to pairwise neuron connectivity: pre-synaptic neuron and post-synaptic neuron couple at the synapse. Our paper describes an interesting way of building DenseAMs using astrocytes, which is a prominent cell type in the glia family. The unique anatomical connectivity of astrocytes, which couple to neurons through tri-partite synapses (pre-, post-synaptic neurons, and the astrocyte’s process) create an elegant theoretical possibility of coupling more than two neurons and mediating the information transmission between those neurons. Key takeaways: 🔷 Astrocytes compute. 🔷 Dense Associative Memories (and other AI architectures) can be built using astrocytes. 🔷 From the computational perspective astrocyte is not just a single entity, rather it is a network of processes. 🔷 Memories can be stored, at least partially, in the biochemical machinery inside the astrocyte, as opposed to just synapses. It was a pleasure to work on this idea together with Leo Kozachkov @Leokoz8 and Jean-Jacques Slotine. Paper: pnas.org/doi/abs/10.107… The image below summarizes the main message of our paper: astrocyte = network of processes that engage in information exchange and computation. The image shows a 3D reconstruction of an astrocyte from the visual cortex using Imaris Bitplane 9.9. A custom Imaris extension, 3D Sholl Analysis, was used to quantify the number of process intersections at 5 μm intervals from the cell soma, marked by the sphere, thereby visualizing the highly complex network of processes of the astrocyte. The raw image was captured in October 2023 using an Olympus FV 3000 confocal microscope with a 60x objective lens and a z-step size of 0.5 μm across a 50–60 μm z-stack for high magnification and resolution. The overlaid network illustrates a mathematical model, which represents the astrocyte as a network of processes that enables heterosynaptic communication and supports high information storage associative memory function. Experimental data credit: Leyka Nagendren @LNagendren , Cagla Eroglu @c_eroglu, Duke University. Artist Credit: Annette Hui, IBM Research, annettehui.com













Excited to share our work on data science in large neuroscience collaboration published in Neuron. We found some challenges but also opportunities on data integration, sharing and research training. Hope it paves the way for further advancing the field. kwnsfk27.r.eu-west-1.awstrack.me/L0/https:%2F%2…