Courtney Shearer
217 posts


Courtney Shearer
@c_sheare
Visiting at GDM Genomics Harvard PhD SSQB @DeboraMarksLab Turtles all the way down



Introducing DroPE: Extending the Context of Pretrained LLMs by Dropping Their Positional Embeddings pub.sakana.ai/DroPE/ We are releasing a new method called DroPE to extend the context length of pretrained LLMs without the massive compute costs usually associated with long-context fine-tuning. The core insight of this work challenges a fundamental assumption in Transformer architecture. We discovered that explicit positional embeddings like RoPE are critical for training convergence but eventually become the primary bottleneck preventing models from generalizing to longer sequences. Our solution is radically simple: We treat positional embeddings as a temporary training scaffold rather than a permanent architectural necessity. Real-world workflows like reviewing massive code diffs or analyzing legal contracts require context windows that break standard pretrained models. While models without positional embeddings (NoPE) generalize better to these unseen lengths, they are notoriously unstable to train from scratch. Here, we achieve the best of both worlds by using embeddings to ensure stability during pretraining and then dropping them to unlock length extrapolation during inference. Our approach unlocks seamless zero-shot context extension without any expensive long-context training. We demonstrated this on a range of off-the-shelf open-source LLMs. In our tests, recalibrating any model with DroPE requires less than 1% of the original pretraining budget, yet it significantly outperforms established methods on challenging benchmarks like LongBench and RULER. We have released the code and the full paper to encourage the community to rethink the role of positional encodings in modern LLMs. Paper: arxiv.org/abs/2512.12167 Code: github.com/SakanaAI/DroPE

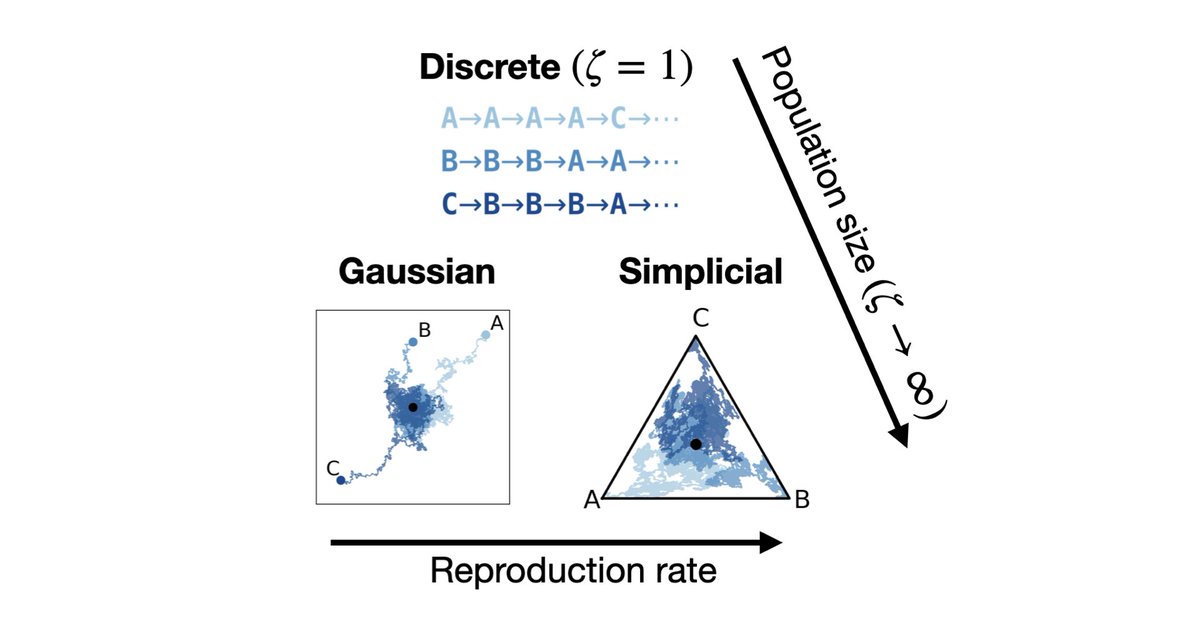
🚨New paper 🚨 Can protein language models help us fight viral outbreaks? Not yet. Here’s why 🧵👇 1/12






What if we have a big protein but need a smaller version for delivery or engineering? Introducing SCISOR: we shrink proteins by training a diffusion model to find natural substrings! arXiv: arxiv.org/abs/2511.07390 @EthanBaron78553, @ruben_weitzman, @deboramarks, @andrewgwils! 1/7

New paper “Proteome-wide model for human disease genetics” is now live at Nature Genetics: rdcu.be/eRu7K popEVE (pop.evemodel.org) finds the needles in the haystacks of human genetic variation:


Super excited about this competition! Use your own developability predictors or build your own on our public dataset of 250 antibodies. We're hoping to benchmark the current state-of-the-art hydrophobicity/thermostability/etc models out there 🏆 (1/2)


Best Paper Award Winners: @setlur_amrith, @AlanNawzadAmin, @wanqiao_xu, @EvZisselman




🚨 New paper 🚨 RNA modeling just got its own Gym! 🏋️ Introducing RNAGym, large-scale benchmarks for RNA fitness and structure prediction. 🧵 1/9