Sabitlenmiş Tweet

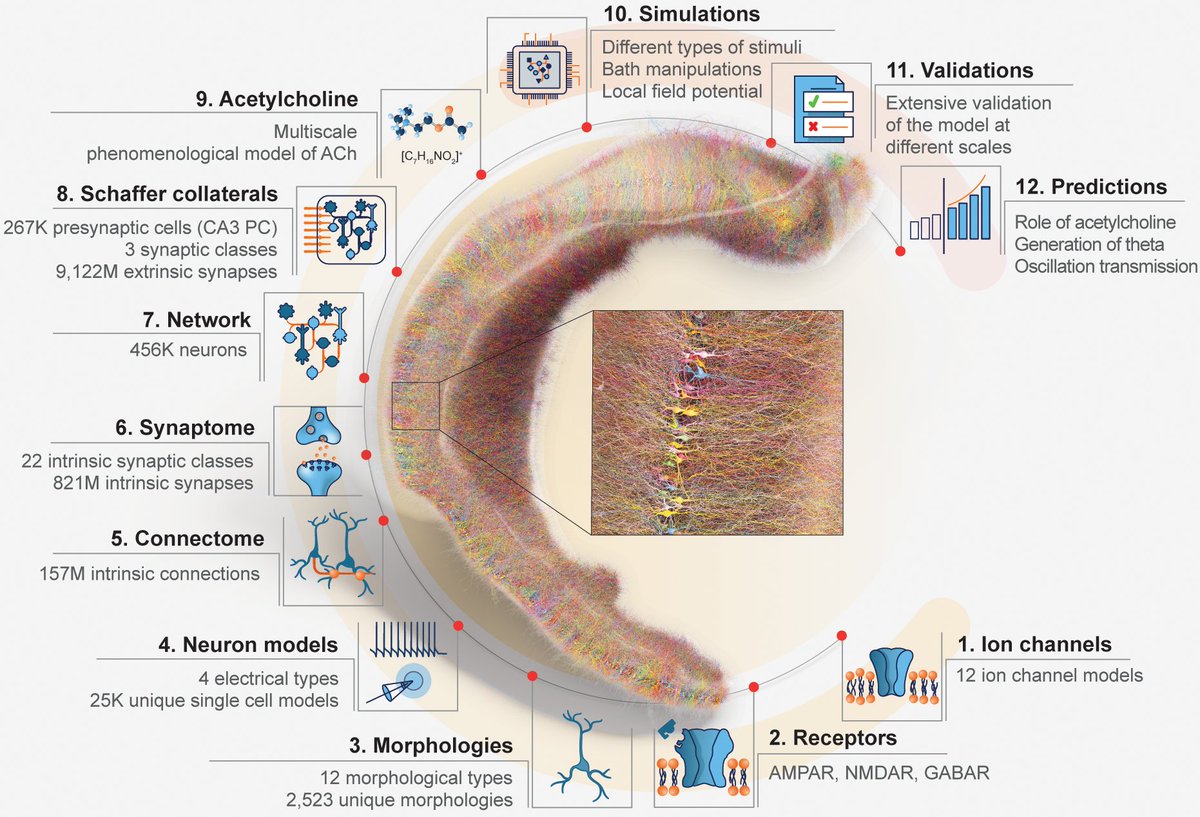
🚨 In our new preprint, and using an organoids model, we show that miRNAs act within a precise temporal window to control cell fate decisions in the human forebrain.🧠
biorxiv.org/content/10.648…
English
Cledi Cerda-Jara
1.3K posts

@cledi_cj
Dr. (rer. nat.) 🧠 Neuronal gene regulation 🔬 Non-coding RNAs and synaptic plasticity 👩🔬Chilean scientist in Germany | She/Her









I’m 25. Give me oddly specific life tips. No general ”surround yourself with positive people” tips. I want the most random, specific advice possible.










Pooled CRISPR screens with joint single-nucleus chromatin accessibility and transcriptome profiling go.nature.com/4hXER5O