
Final #ASHG24 poster session is starting now, come say hi if you’re interested in long read PGx, poster 7021!
Chris Saunders
293 posts

@ctsa11
Rare disease and cancer analysis models for sequencing data. Scientist at PacBio. Art school survivor. Views my own. https://t.co/5iXEF3EMSK

Final #ASHG24 poster session is starting now, come say hi if you’re interested in long read PGx, poster 7021!



We assessed our new method on the recent draft GIAB SV benchmark from the T2T-HG002-Q100 assembly, finding sawfish has the highest F1-score among evaluated callers on every tested SV size range and depth level. Other callers required at least 30x depth to reach sawfish at 15x:



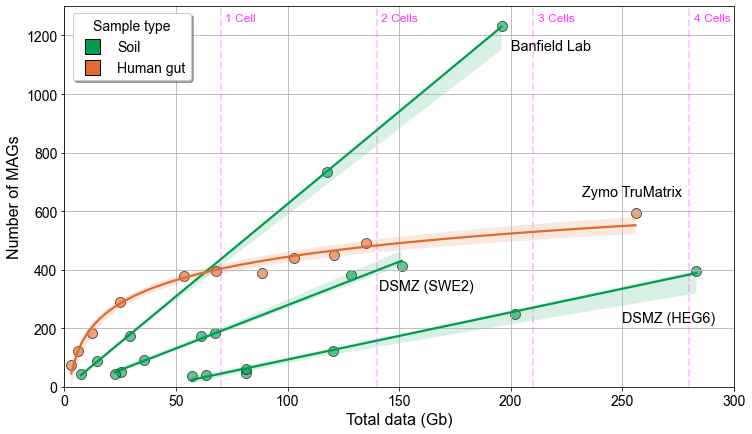

Unlock the complexities of soil communities with #HiFisequencing on #Revio. Discover how to obtain high-quality MAGs with more data and lower costs than ever before. Register now: bit.ly/3yVbFuG #PacBio

Happy to introduce sawfish, a new HiFi structural variant caller emphasizing local haplotype modeling to improve SV representation and genotyping in both single and joint-sample analysis contexts: 1/n doi.org/10.1101/2024.0…

Local assembly of @PacBio HiFi for exceptionally accurate structural variant detection. Work by @ctsa11 @holtjma @nothingclever @dnb_hopkins @MikeEberle software: github.com/PacificBioscie… Preprint: doi.org/10.1101/2024.0…

