Brandon White@bwhite5290
To replace animal testing with AI, we need MASSIVE human datasets.
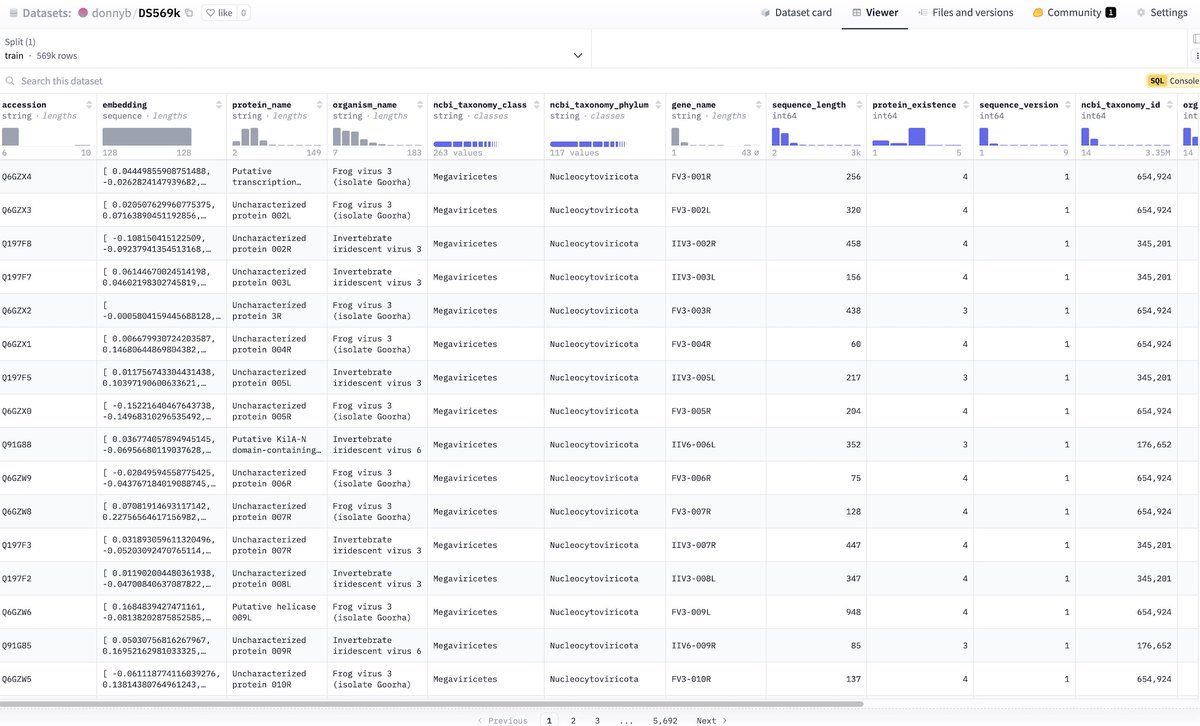
Today, we're thrilled to share Axiom's new data exploration tool, providing the ability to visually explore the world's largest primary human liver toxicity dataset. Built with Axiom's proprietary wetlab protocols, our dataset includes detailed liver toxicity profiles for over 100,000 distinct molecules.
The key to this dataset is our ability to do high-throughput, multiplexed high-content screening with primary human liver cells. Traditionally, toxicity assays either sacrifice throughput or sacrifice biological relevance (using easy-to-grow immortalized cell lines instead of real human cells). We managed to combine throughput, physiological relevance, and multiplexing in one platform. The assays run in a high throughput format using automation, meaning thousands of compound-dose conditions can be tested in one experiment. We achieved this using pooled primary human hepatocytes, which are often fragile and expensive. By systemizing our automation and quality control processes, we were able to run over 120+ batches on the same donor pool with incredible reproducibility and consistency.
We did this while integrating many readouts per well, whereas many existing toxicity assays only do a single readout. Our multiplexed approach provides far more data per experiment enabling us to measure 10-20 different toxicity phenotypes such as apoptosis, necrosis, mitochondrial fission, endoplasmic reticulum stress, stress granule formation, microtubules, and more all from a single well on a 384-well plate! The combination of scale, high content information, and data quality is exactly what is needed to train highly accurate AI models in biology.
If you're interested, please explore the dataset in the comments below and let me know if you want to chat about the details!