
Florian
250 posts
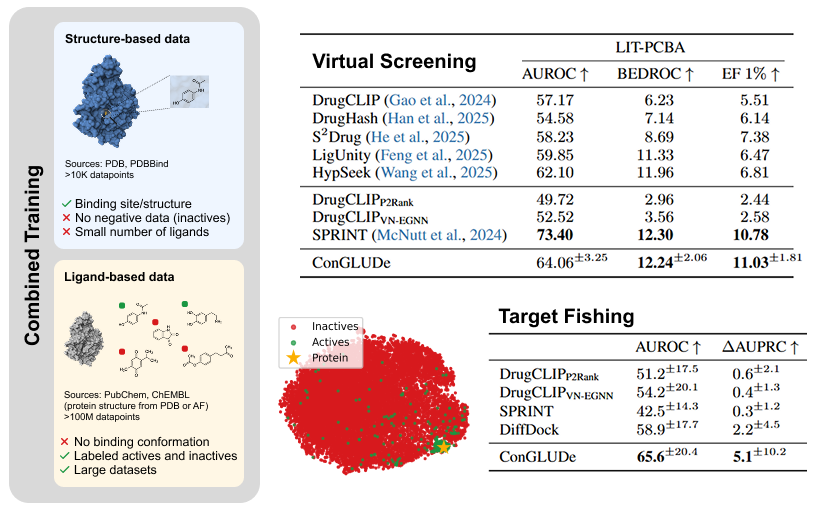






BIG BREAKTHOUGH: A new AI tool could dramatically speed up the discovery of life saving medicines. Researchers at Tsinghua University created a new system called DrugCLIP, that can screen drug molecules against human proteins at a speed that makes traditional methods look ancient. > DrugCLIP uses deep contrastive learning to turn both molecules and protein binding pockets into vectors and match them almost instantly. > It screened 500 million molecules across 10,000 human proteins, covering half of the entire human druggable proteome. > The system completed 10 trillion molecule protein evaluations in a single day, roughly 10 million times faster than classic docking simulations. > They used AlphaFold2 to generate protein structures and then refined binding pockets with a custom tool called GenPack. > The model even identified compounds for TRIP12, a protein linked to cancer and autism that has resisted traditional drug-targeting approaches. All data and models are open access, so labs worldwide can now speed up early stage drug discovery.







BERT is just a Single Text Diffusion Step! (1/n) When I first read about language diffusion models, I was surprised to find that their training objective was just a generalization of masked language modeling (MLM), something we’ve been doing since BERT from 2018. The first thought I had was, “can we finetune a BERT-like model to do text generation?”










Perceiver IO is good reading/pointers for neural net architectures arxiv.org/abs/2107.14795 esp w.r.t. encoding/decoding schemes of various modalities to normalize them to & from Transformer-amenable latent space (a not-too-large set of vectors), where the bulk of compute happens.

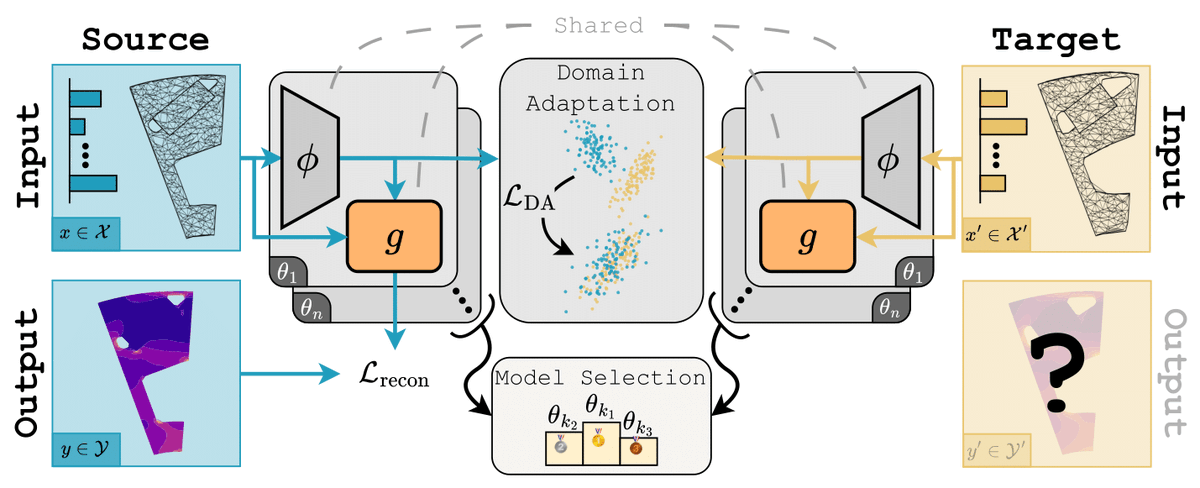

After a long hiatus, I've started blogging again! My first post was a difficult one to write, because I don't want to keep repeating what's already in papers. I tried to give some nuanced and (hopefully) fresh takes on equivariance and geometry in molecular modelling.