GPTomics retweetledi

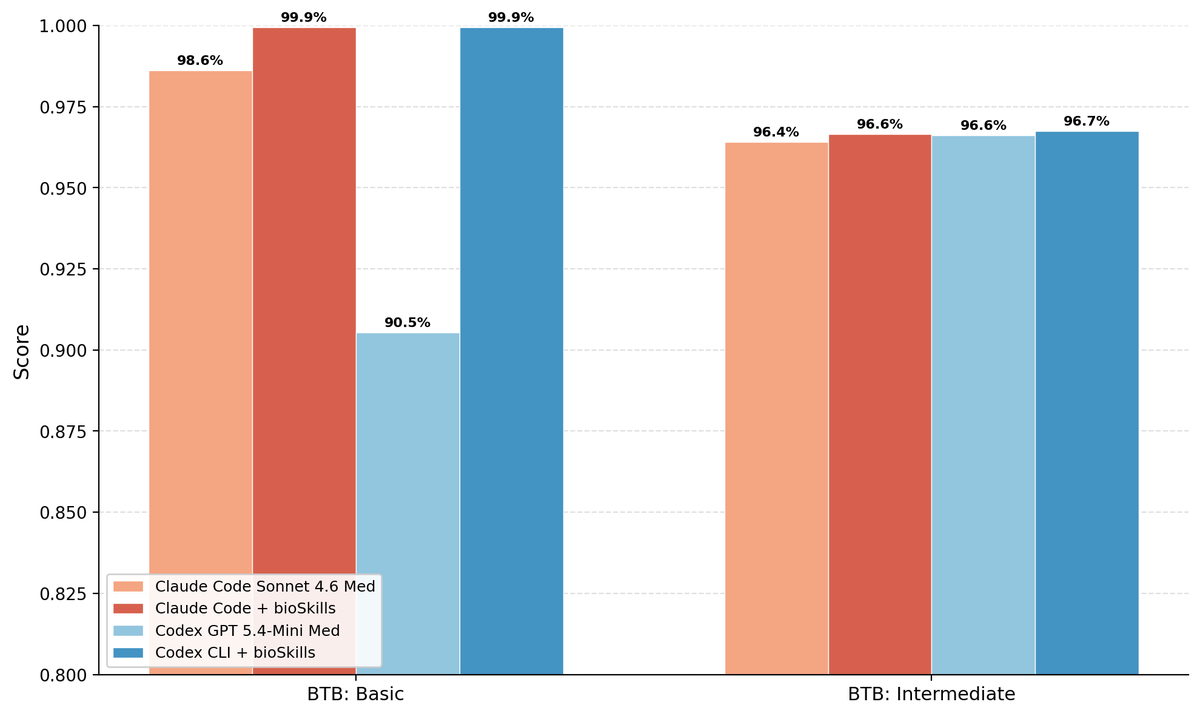
Took this for a spin on both @claudeai Code and @OpenAI Codex both with and without BioSkills. Seeing both some excellent performance without the skills and some clear skills based improvement
GPTomics@gptomics
Created a new set of 33 tasks and a harness to benchmark agents on bioinformatics datasets. github.com/GPTomics/bioTa… Take it for a spin
English