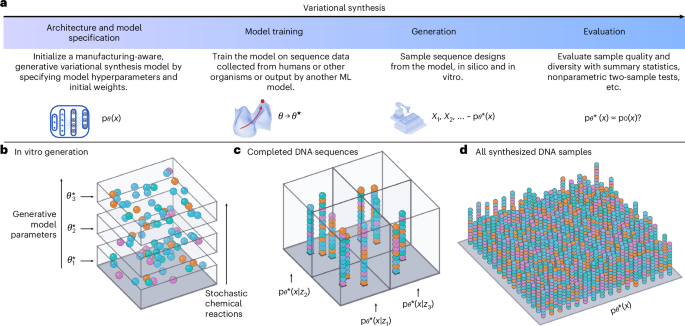
How much design and screening do you need? Amazon had to design 288,000 nanobody constructs spanning eight target epitope regions and screened 100,000 using yeast display. Thats the scale still required for de novo design amazon.science/publications/a…