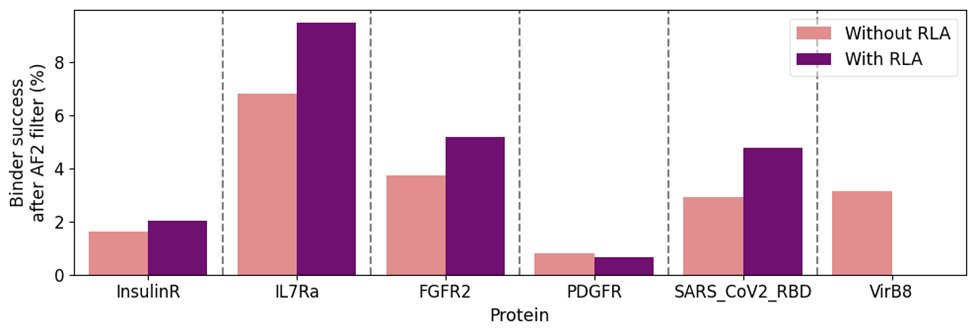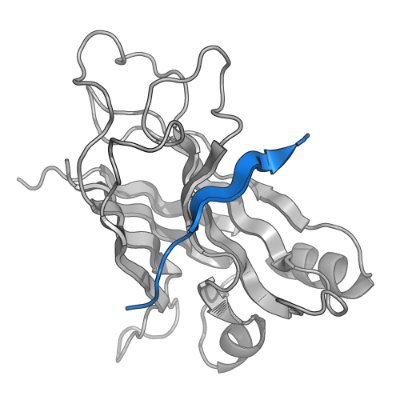
To solve this, Birnbaum and Keating present PottsMPNN, a sequence design model that learns a sequence-energy landscape from MSAs. PottsMPNN outperforms other sequence design models and is a drop-in alternative to ProteinMPNN. Code: github.com/KeatingLab/Pot…
(2/2)
English