Tobias Kroniger retweetledi
tidyplots
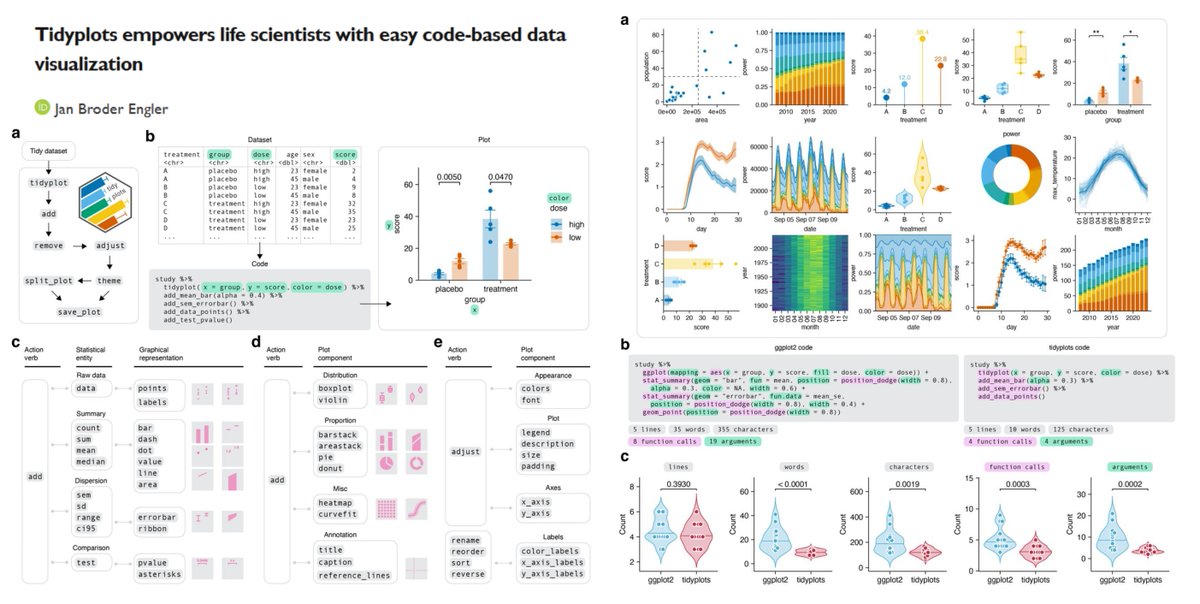
Time to say goodbye to ggplot2?🫡
"a significant reduction of code complexity" vs ggplot2
cran.r-project.org/web/packages/t…
@JanBroderEngler bioRxiv 2024
biorxiv.org/content/10.110…
English
Tobias Kroniger
82 posts


@kronigert
Field Application Scientist @ Bruker Daltonics. Views are my own. he/him. #Proteomics #TeamMassSpec






76/432 samples in on my big TimsTOF run and looking solid 🤞 Amazing work from our new MSc and PhD students. Going to be gritting my teeth all weekend.... #TeamMassSpec #Proteomics
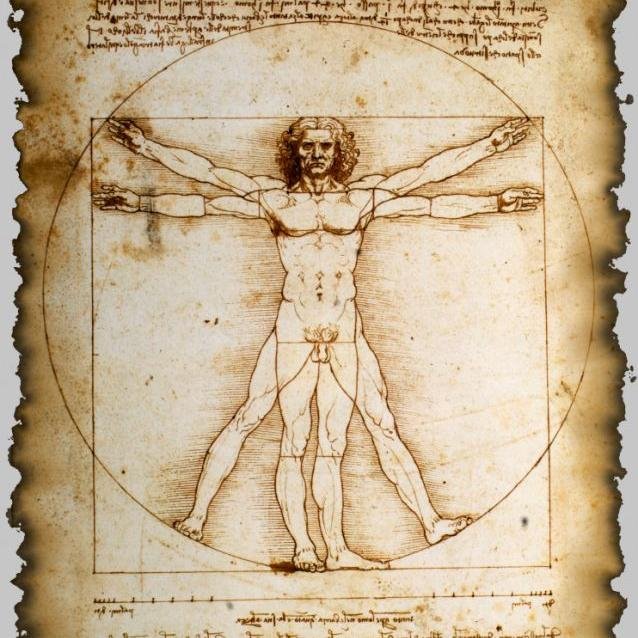





High throughput single-cell proteomics of in-vivo cells biorxiv.org/content/10.110… --- #proteomics #prot-preprint


















