Laboratory of biomolecular modelling and design
308 posts


Laboratory of biomolecular modelling and design
@kullbmd
Arnout Voet, Computational protein and drug design, structural biology, biomolecular modelling, protein crystallography and biophysics.




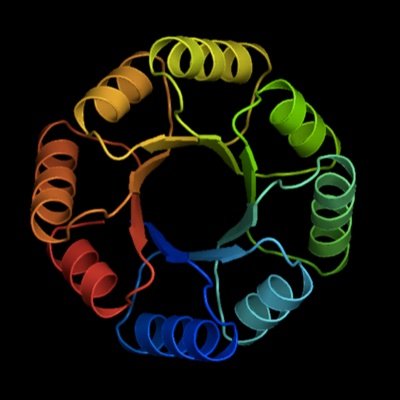

Sparks of function by de novo protein design go.nature.com/48jXz1z



BREAKING NEWS The Royal Swedish Academy of Sciences has decided to award the 2024 #NobelPrize in Chemistry with one half to David Baker “for computational protein design” and the other half jointly to Demis Hassabis and John M. Jumper “for protein structure prediction.”






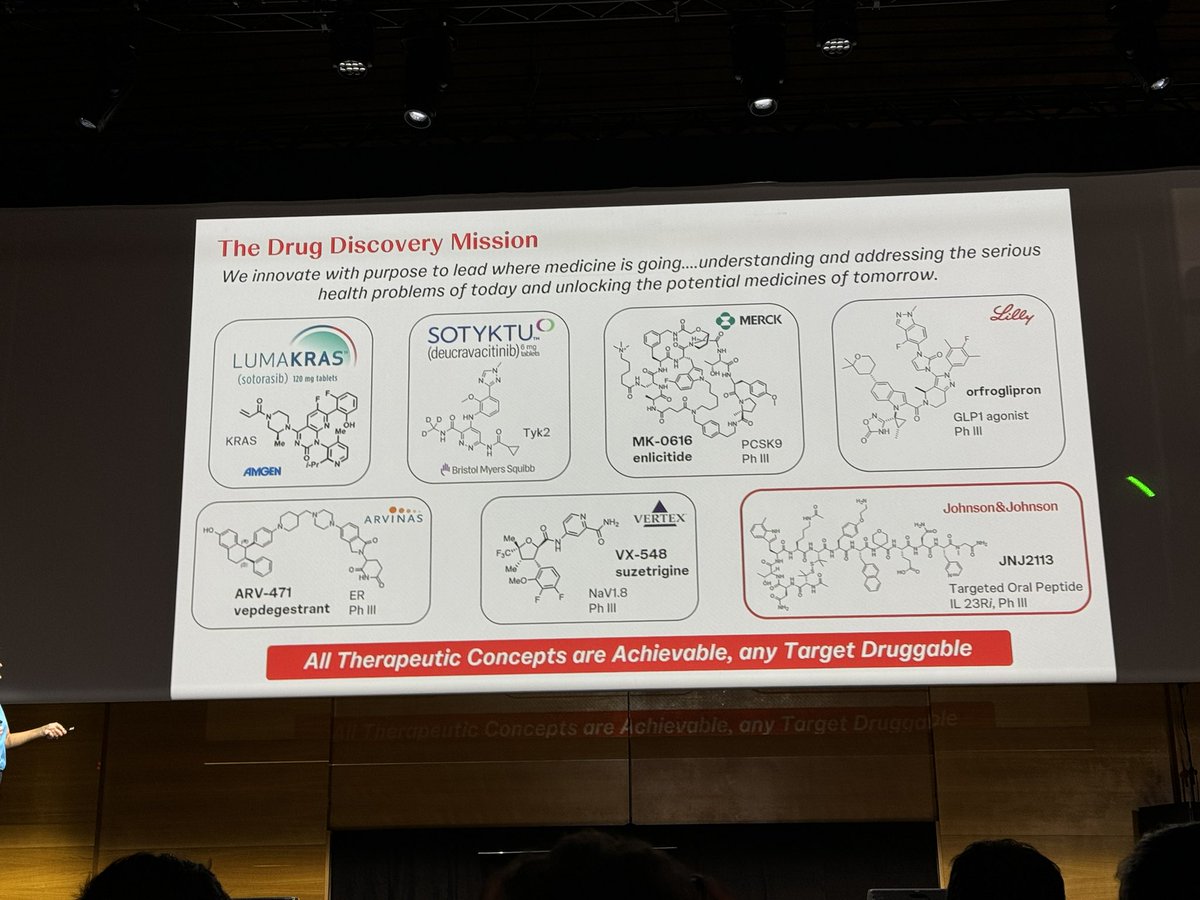
JJ’s Emma Parmee with a tour de force about chemistry success factors in drug discovery at #EFMCISMC24: Mastering structural complexity to drug more complex targets, the importance of hit ID diversity, HTE to speed up Make cycles, D2B for #PROTAC hit ID, Surf-DELs @EuroMedChem


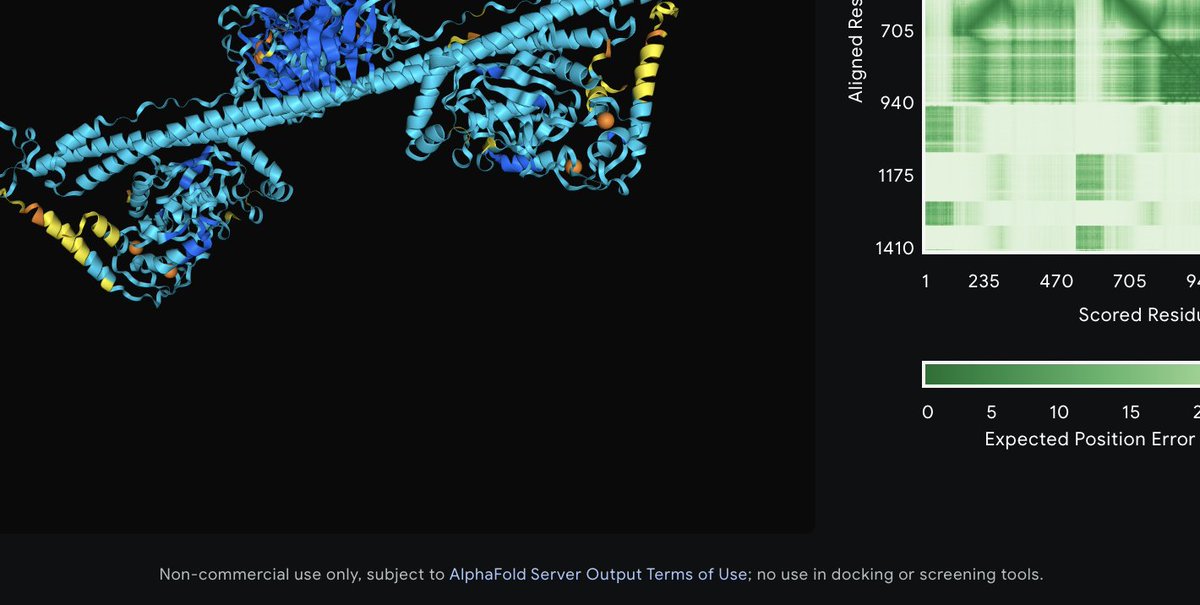
The worst thing about AlphaFold3 (after no code) is that you can't use a protein structure prediction to do drug development (if I read this correctly). @GoogleDeepMind must own all drugs developed from AF3. It's basically The Matrix for protein structure prediction.

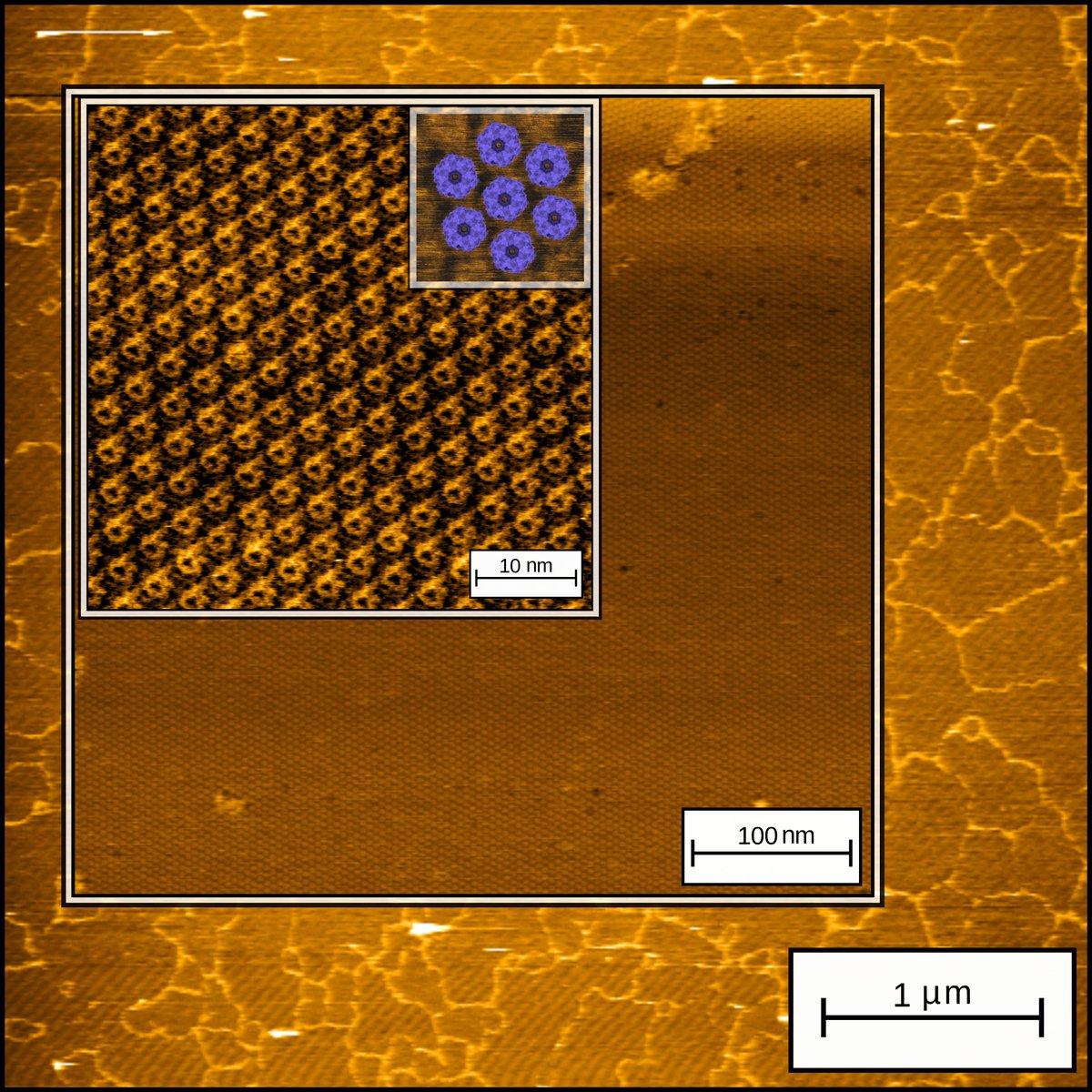
Sometimes #AFM gives you this joy! micrometer wide Protein-based 2D layers made from a Computational Designer protein. Excited to see where this protein will lead us next! #BioAFM #AtomicForceMicroscopy #CPD #proteindesign #Nanobiotechnology @kullbmd @KU_Leuven #biochemistry