Andras Lasso
190 posts

Andras Lasso
@lassoan
Interventional medical device R&D lead engineer, researcher, @3DSlicerApp core contributor.
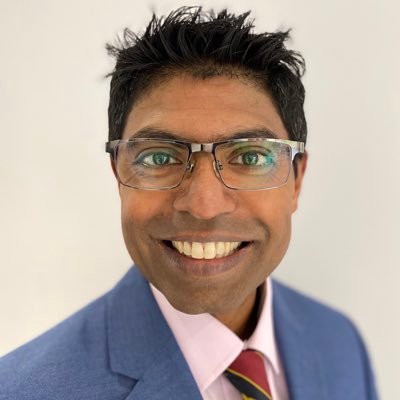






















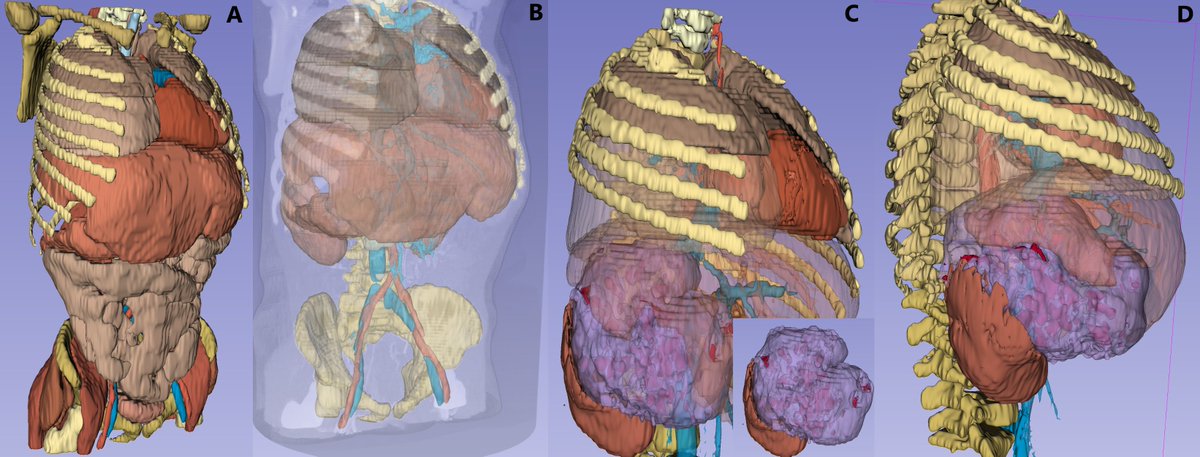










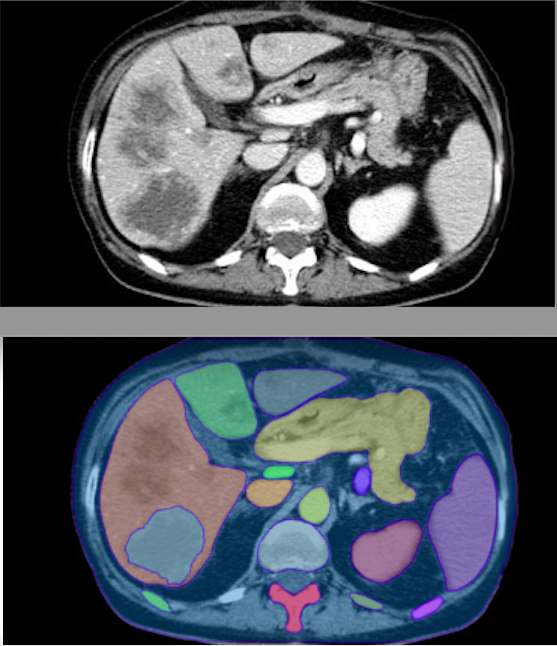
This is a game changer. Segment 100+ structures in any whole-body CT image in 2 minutes using TotalSegmentator in @3dslicerapp All free, open-source software. Runs on any computer, no GPU is required. See more information at discourse.slicer.org/t/26710

Today we're releasing the Segment Anything Model (SAM) — a step toward the first foundation model for image segmentation. SAM is capable of one-click segmentation of any object from any photo or video + zero-shot transfer to other segmentation tasks ➡️ bit.ly/433YuBI







4/5) The multimodality-capable (3DE, CT, CMR) tricuspid valve modeling tools we created for this study are now available open-source @3DSlicerApp #SlicerHeart #SlicerSALT