




Linda B. Baughn
431 posts


@lbaughn
Geneticist and research scientist at Mayo Clinic working to better understand, diagnose and treat B-cell malignancies. Opinions on Twitter are my own.






Dearest gentle reader, we are delighted to announce a new story from our lab published in @Nature describing how a meal's systemic metabolic changes are interpreted by your immune system to enhance adaptive immunity. A thread 1/ nature.com/articles/s4158…


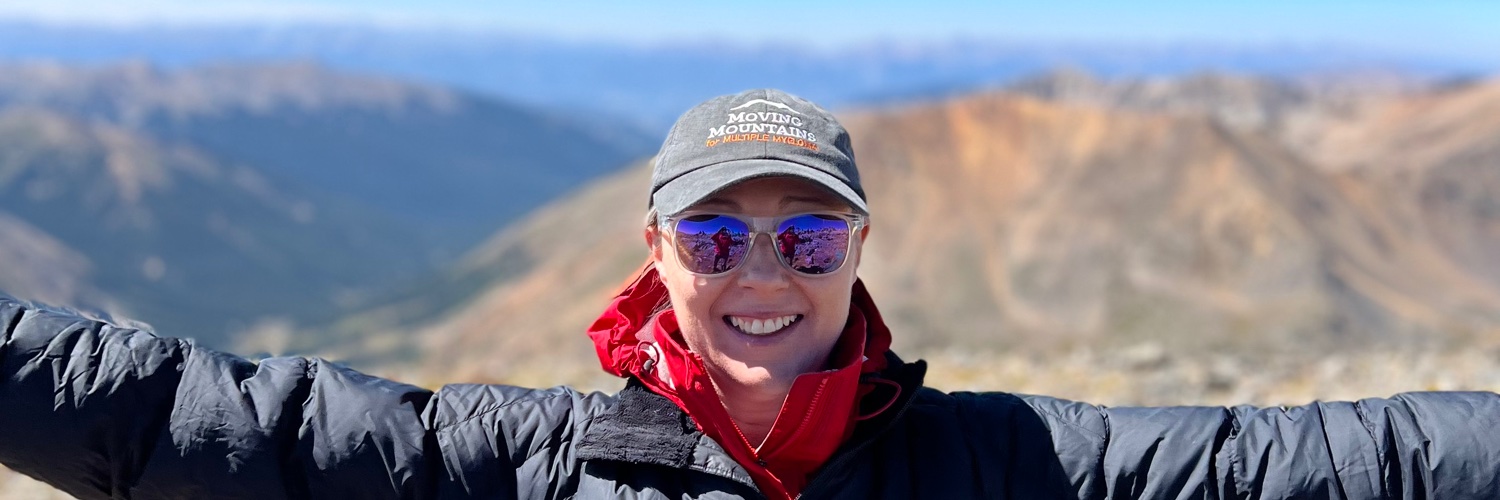
Excited to share our first 2026 paper from the lab! In a collaboration with @MayoClinic @MSKCancerCenter @EmoryUniversity @SylvesterCancer @OsuChsd & @MSK_DeptOfMed, we investigated the drivers of venetoclax resistance in t(11;14) Multiple Myeloma #mmsm. ashpublications.org/blood/article-…



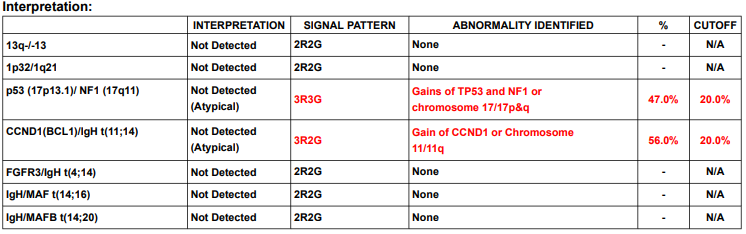



Just out: Guidelines for testing and reporting cytogenetic results in myeloma @BloodCancerJnl #OpenAccess Bookmark! Allows you to incorporate latest IMS/IMWG high risk criteria in practice. @lbaughn @RahulBanerjeeMD @Rfonsi1 nature.com/articles/s4140…



#Myeloma Paper of the Day: Eagerly awaited new high-risk IMWG definition: 1. del 17p w/ >20% clonal frac &/or TP53 mut; 2. t(4;14), t(14;16), t(14;20) w/ 1q+ &/or del 1p32; 3. monoallelic del 1p32 w/ 1q+ or biallelic del 1p32; 4. β2 ≥5.5 & nl SCr: pubmed.ncbi.nlm.nih.gov/40489728/. #mmsm









Don’t say “atypical abnormal” unless it is - @RahulBanerjeeMD @lbaughn @VincentRK @Rfonsi1 oncodaily.com/blog/154062 #Cancer #CancerAwareness #CancerCare #IMS2024 #IMS24 #OncoDaily #Oncology #Medicine #Health #MedEd #MedX #MedNews

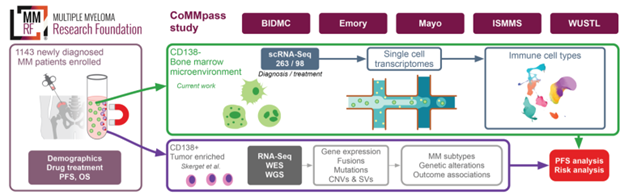


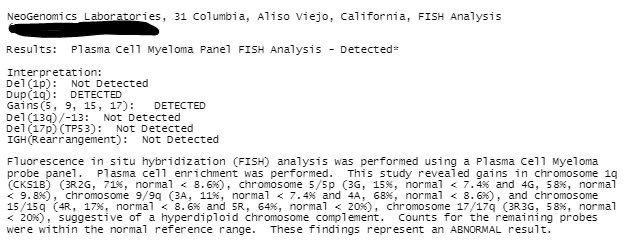
#MMsm and it happens again - a patient diagnosed with “high-risk myeloma” almost a year ago whom I’ll undiagnose this month. 🚨 Only call it del(17p) if you see the whites of its eyes in the path report. Thanks to @lbaughn et al who are working on changing this!




We have to reach physicians in the community to see how we can get their input. Can @ACCCBuzz help to get this survey out and partnering with them. Apart from risk stratification, can we report that BCL2 inhibitors can be used in the FISH report. @PedalheadPHX @lbaughn


Thank you to those who have completed the myeloma FISH survey! We are so grateful. If you haven’t yet completed the survey, we ask you to please consider making your voice heard. Your responses are so valuable. #MMSM @RahulBanerjeeMD