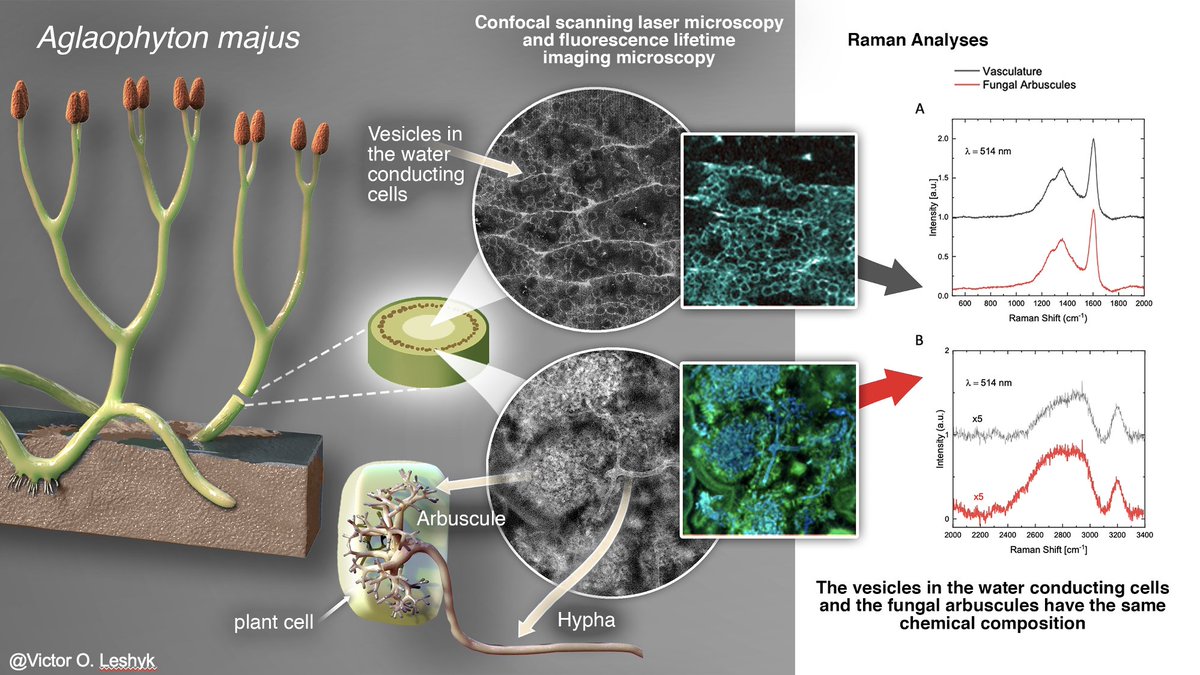
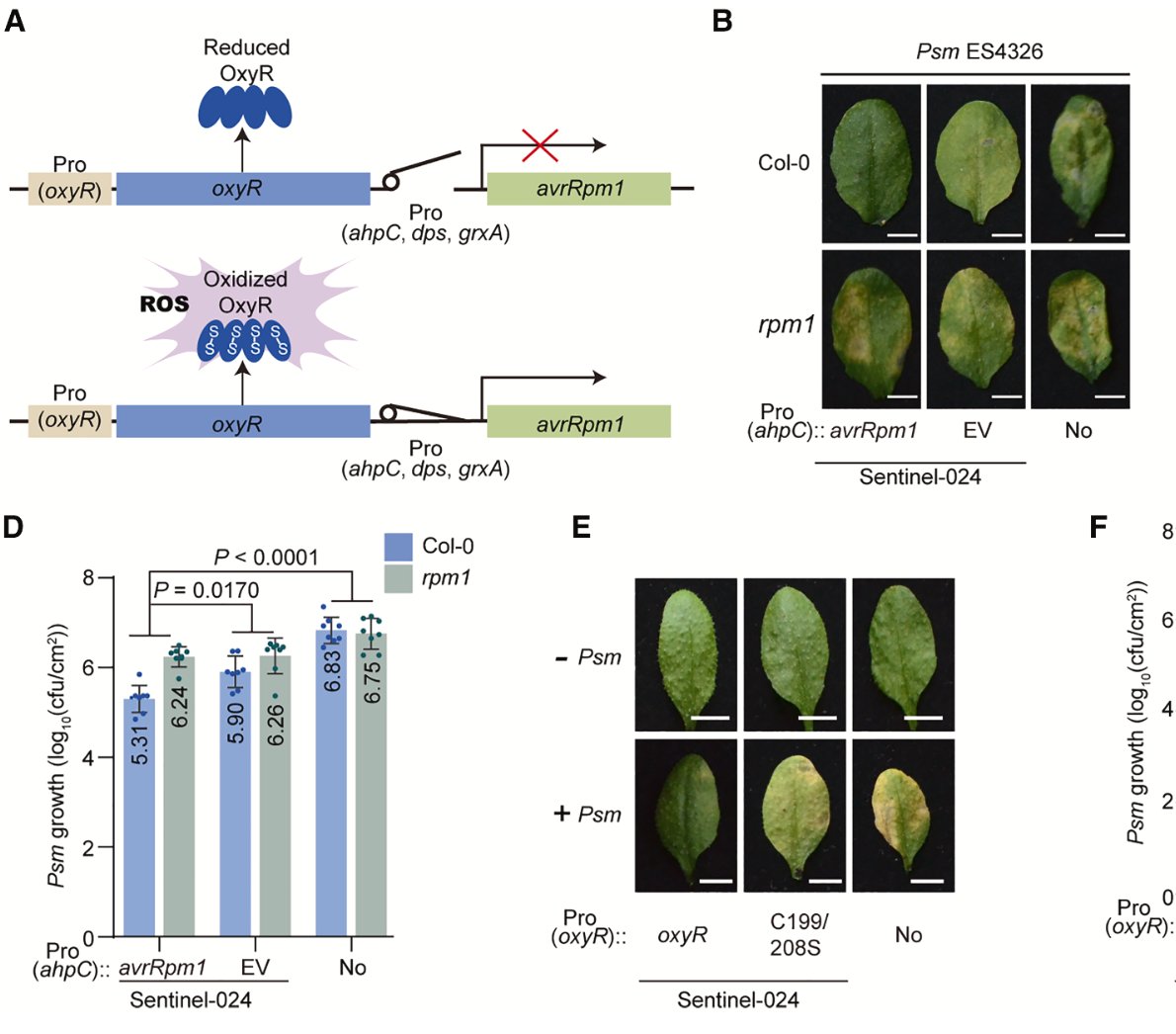
1/ It is my pleasure to share the latest preprint of the team: "Symbiotic diversification relies on an ancestral gene network in plants" doi.org/10.1101/2025.0… Here, we identified and functionally validated a novel master regulator of intracellular symbioses! A thread ...