
Summer course 4-8 August 2025 at Aarhus University, with professor Luc Janss @lljanss as lecturer🌞: #GenomicPrediction in #Animals and #Plants Read more here 👉international.au.dk/education/admi… Application deadline is 18 March 2025❗
Luc Janss
32 posts

@lljanss
Professor, plant genetics and breeding, big data, prediction, Aarhus University, Denmark.

Summer course 4-8 August 2025 at Aarhus University, with professor Luc Janss @lljanss as lecturer🌞: #GenomicPrediction in #Animals and #Plants Read more here 👉international.au.dk/education/admi… Application deadline is 18 March 2025❗





Global diversity in faba bean: Identification of QTLs for key agronomic traits and genomic signatures of selection @stiguandersen rdcu.be/danWg



Marking a milestone in #plant #genomics!!! An amazing team of scientists around the globe unraveled the giant #fababean genome (~ 13 Gb), which is now out @Nature ..Huge congrats to all authors!!! #plantprotein @LeibnizIPK @LeibnizWGL nature.com/articles/s4158…
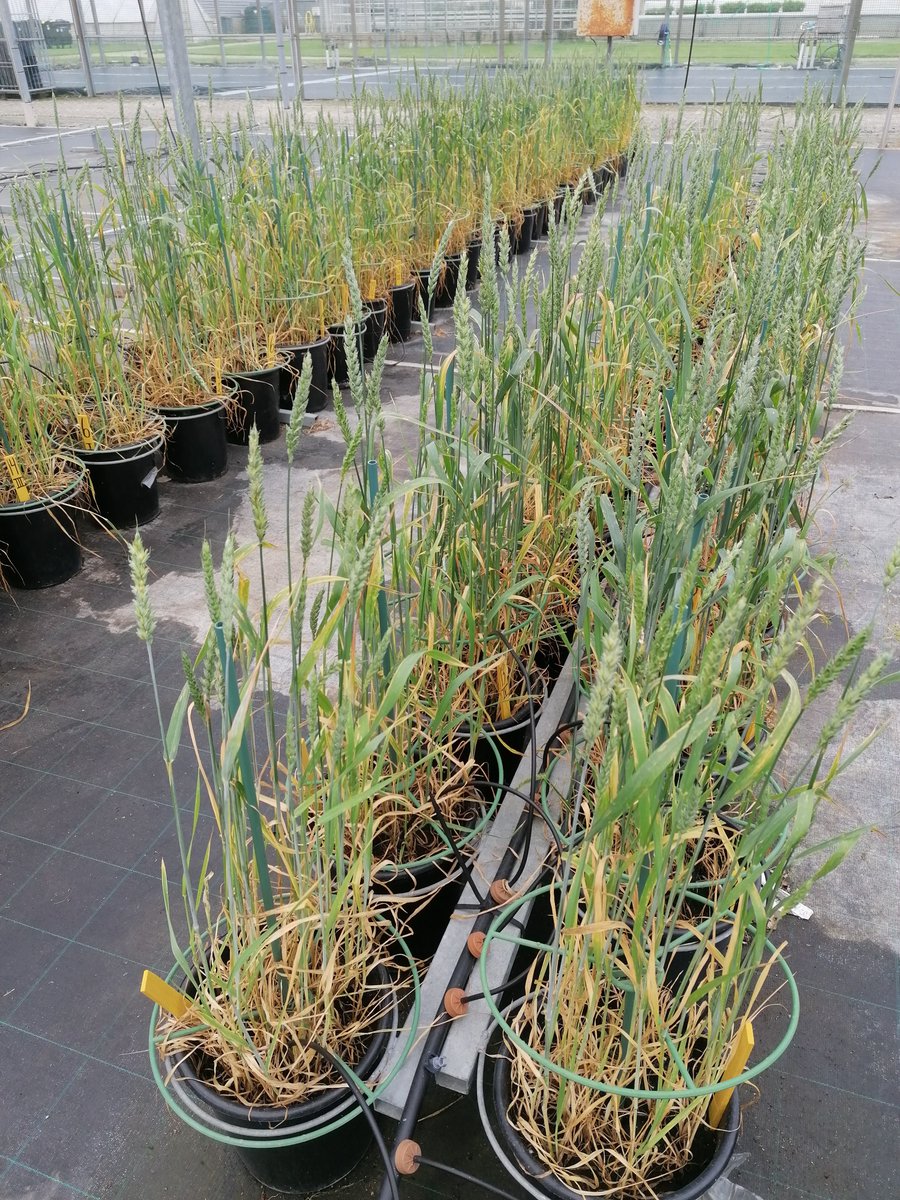




Our latest results on genetics of white clover yield based on continuously tracking growth 3600 plants are now published in @TheorApplGenet rdcu.be/czgaX #legumes #plantgenetics @smoeskjaer @lljanss @DLF_Seeds


A big step towards faba beans free from vicine and convicine. @GeuFloresF @alanhschulman @fred_stoddard @D_O_Sullivan @NaturePlants #fababean #favism #legumes doi.org/10.1038/s41477…







Congratulations to Pernille Bjarup Hansen who successfully defended her PhD thesis Friday on #Genetic Dissection and #Prediction of Complex #Traits in #Barley and Festulolium Grasses. #QGG_AU bit.ly/2G399Hg @lljanss #Torben_Asp
