Sabitlenmiş Tweet

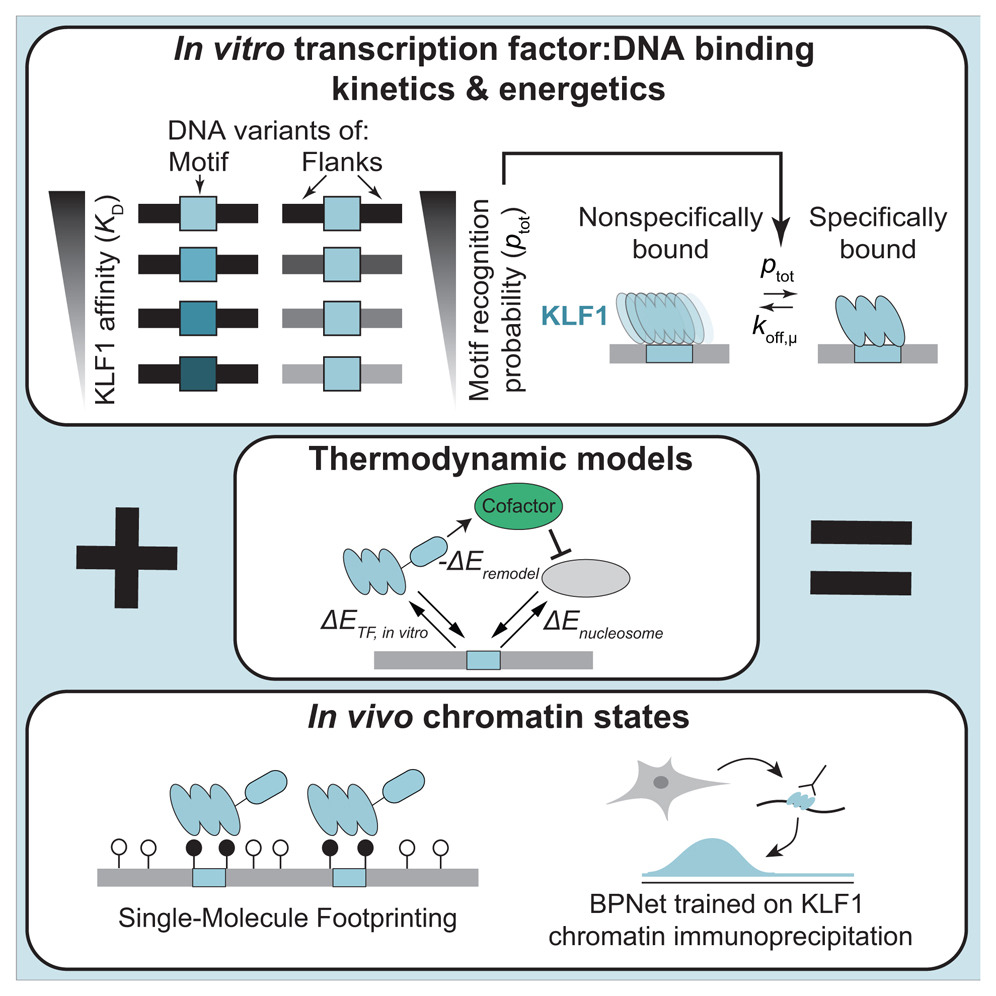
📣 I am happy to announce that our work linking DNA binding affinities and kinetics 𝘪𝘯 𝘷𝘪𝘵𝘳𝘰 and 𝘪𝘯 𝘷𝘪𝘷𝘰 for the human transcription factor KLF1 just got published in @CellCellPress:
cell.com/cell/fulltext/…
(1/2)
English
Emil Marklund
134 posts

@marklundem
Assistant Professor at @scilifelab and @Stockholm_Uni. Intrigued by what makes life work at the molecular level.




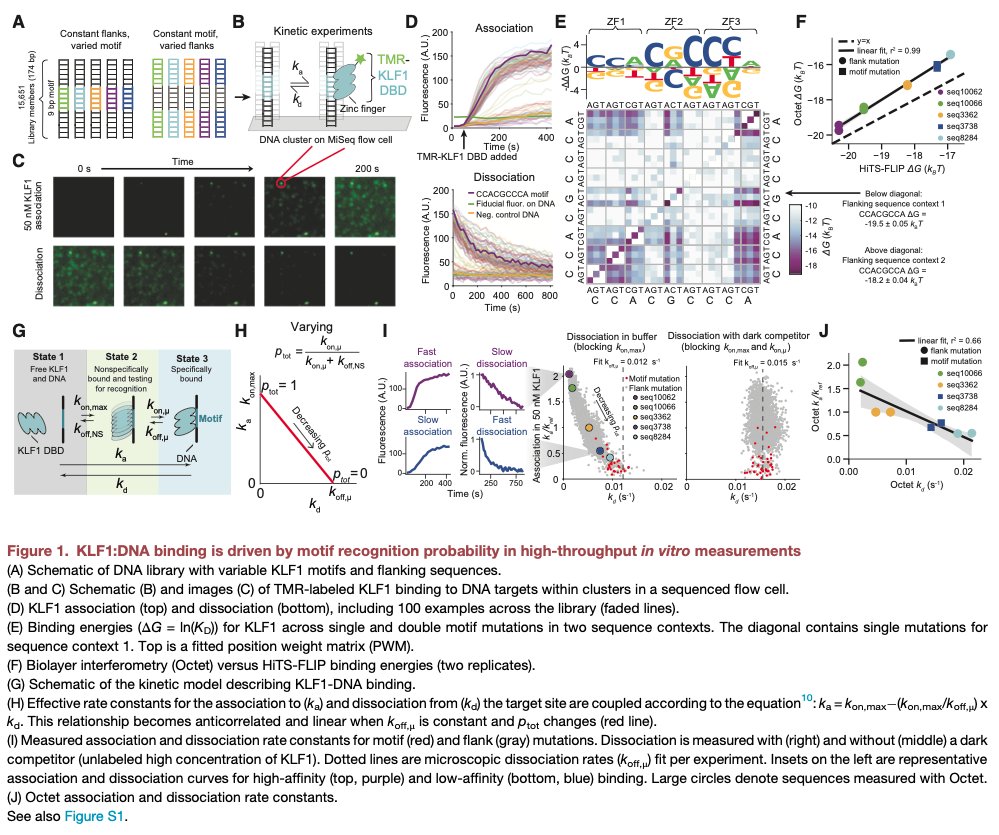


Our work using thermodynamic principles to link in vitro TF affinities and kinetics to single-molecule chromatin states in cells is now on bioRxiv! Amazing effort by first author @juliaschaepe in @WJGreenleaf lab: biorxiv.org/content/10.110… @Stanford @scilifelab @Stockholm_Uni [1/9]









Incredibly excited that our paper linking single molecule states of TF binding to gene expression using quantitative thermodynamic models is out in Nature today. An amazing collaboration with the Bintu Lab. Congrats to Ben, Michaela, and Julia! nature.com/articles/s4158…