Sabitlenmiş Tweet

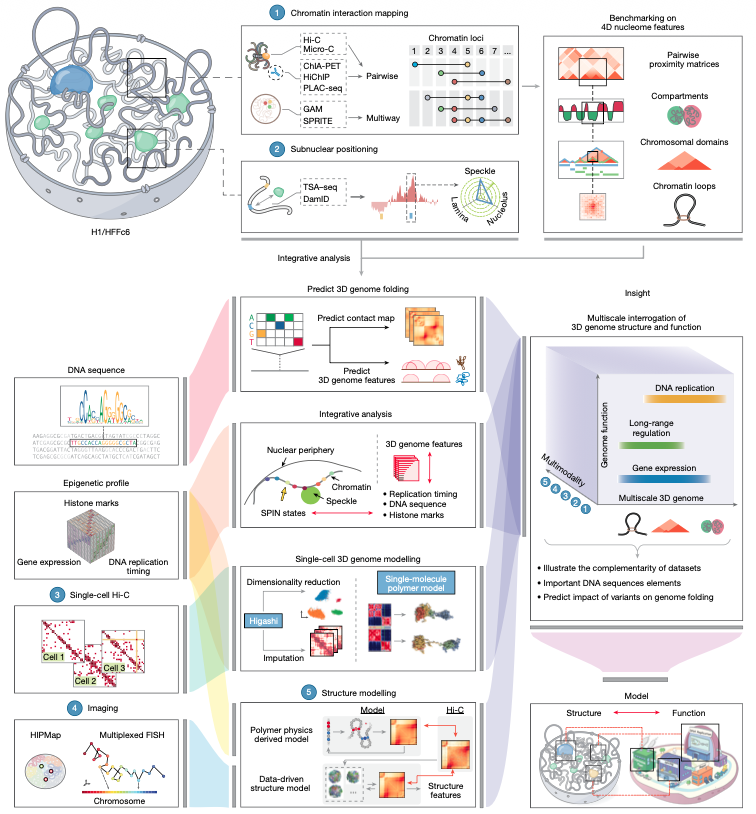
Im so happy to share this part of my PhD thesis just right now in @NatureComms. Check it out! Implications of the 3D chromatin organization for genome evolution in a fungal plant pathogen Special thanks to @MFSeidl and @Team_Thomma nature.com/articles/s4146…
English