PythoNeuro
219 posts

PythoNeuro
@pythoneuro
Promoting python as the best language for coding and analyzing in the neuroscience field! 🧠🧑🎓👩💻 Also love open science ❤️

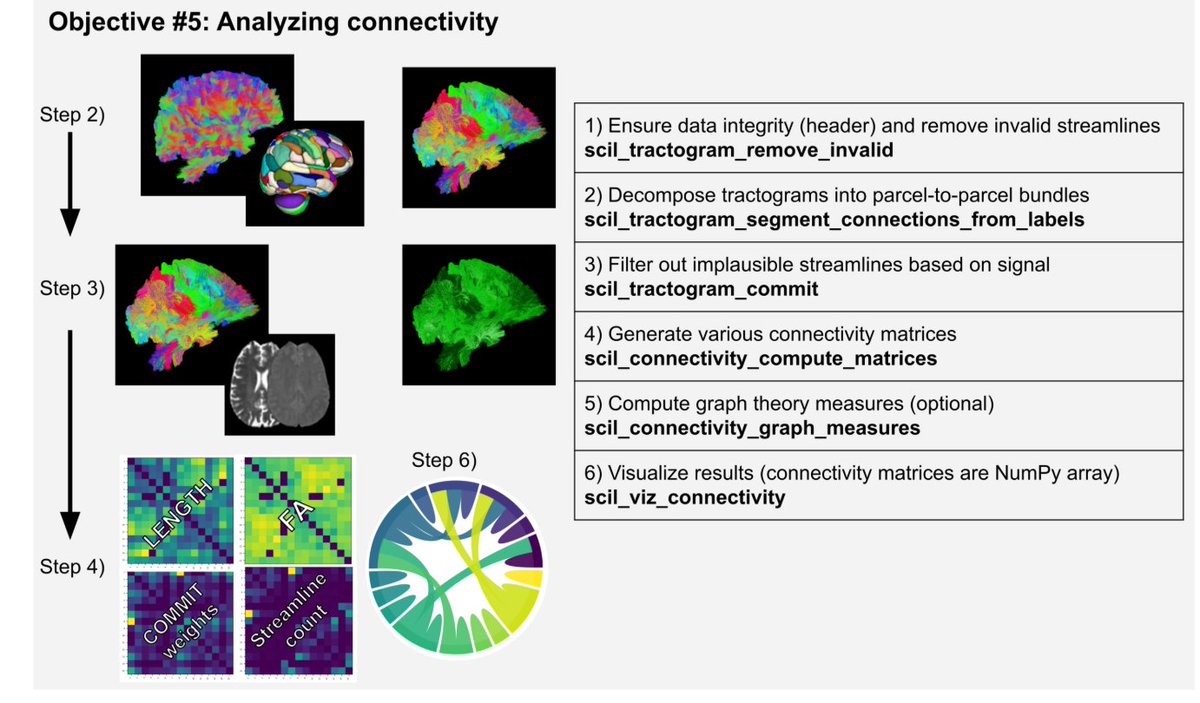
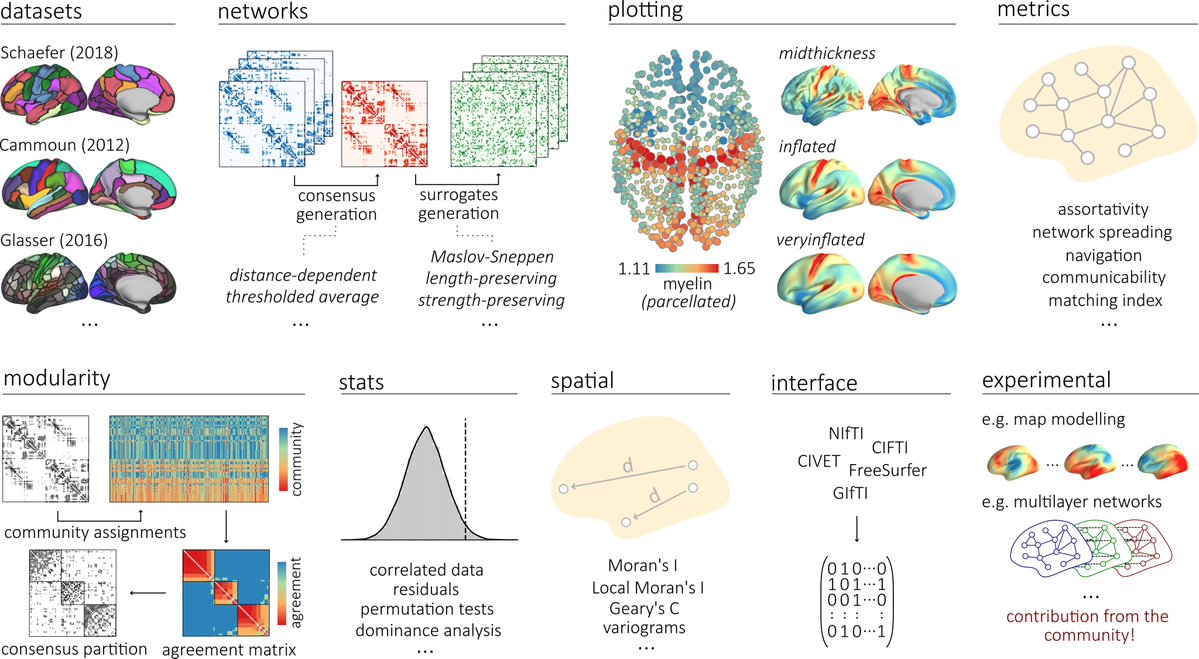
Renauld et al. present scilpy, an open-source #Python library for diffusion magnetic resonance imaging and tractography: doi.org/10.52294/001c.… @OHBM



Visualize genomic data with ease using gggenomes, an R package that extends ggplot2 to handle and display genomic information intuitively. Whether you’re comparing genomes, analyzing features, or showcasing synteny, gggenomes provides the tools you need to turn complex genomic data into clear, informative visualizations. Why use gggenomes? ✔️ Genomic-focused visualizations: Specifically designed for handling genomic data, including features, alignments, and comparative analysis. ✔️ Versatile and modular: Create detailed and layered plots for diverse genomic scenarios with flexibility for customization. ✔️ Built on ggplot2: Leverages ggplot2’s familiar framework, making it easy for users to adapt and enhance their visualizations. The example visualization shown here is taken directly from the gggenomes GitHub repository, demonstrating how it transforms genomic data into compelling plots: github.com/thackl/gggenom… Curious to learn more about creating data visualizations in R and using tools like ggplot2 and its extensions? Check out my online course, "Data Visualization in R Using ggplot2 & Friends!" Further details: statisticsglobe.com/online-course-… #tidyverse #statisticsclass #RStats #database #datavis #R #ggplot2 #Python #VisualAnalytics