Randolph Lopez B.
3.3K posts


Randolph Lopez B.
@randolph380
Co-Founder/CTO of A-Alpha Bio (SynBio+ML+DrugDev). Passionate about world affairs, running, the outdoors and ramen.


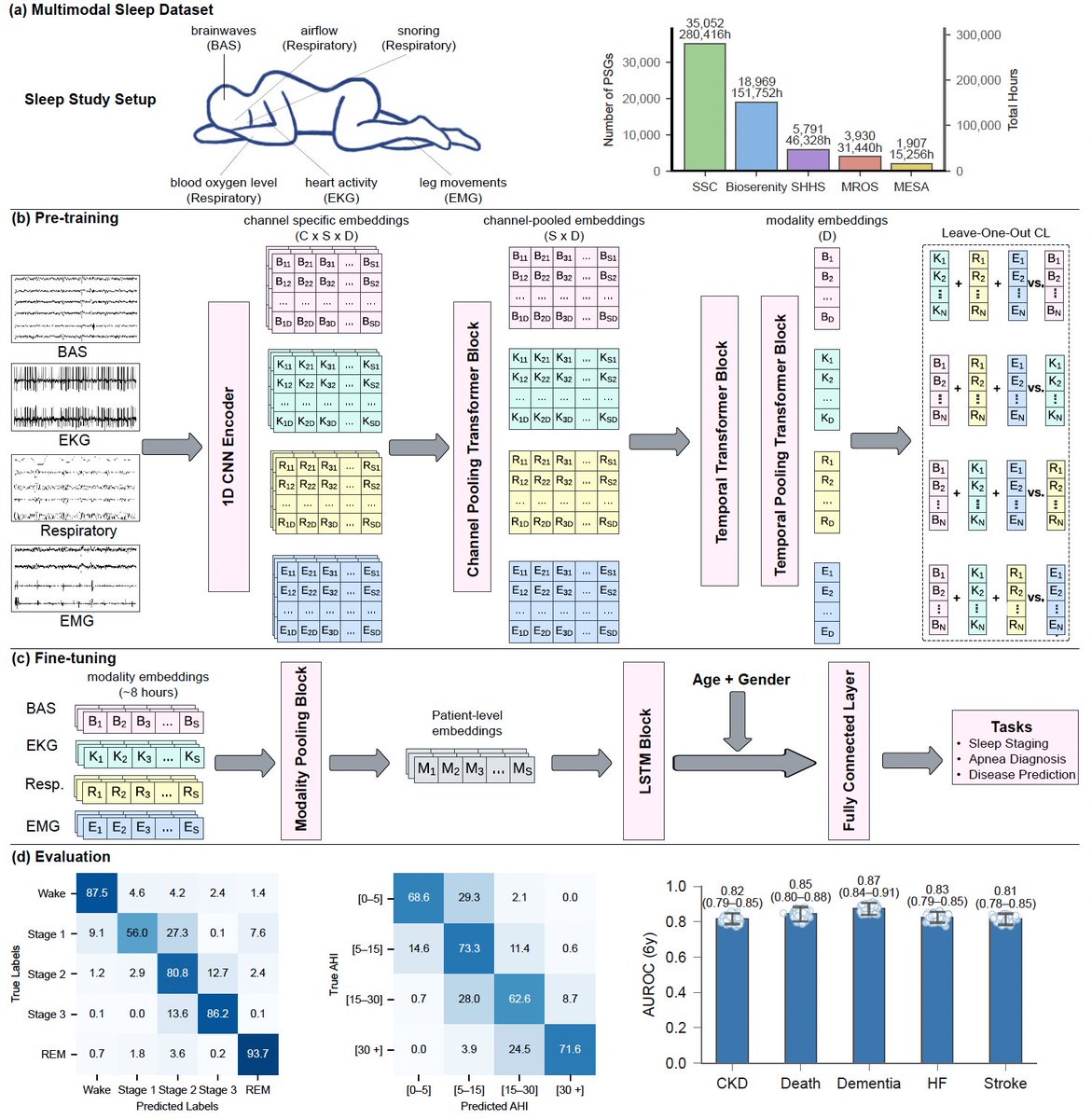

@AAlphaBio paper on their protein-protein interaction model is out! paper: biorxiv.org/content/10.110… for context, i wrote about them a few months ago (owlposting.com/p/creating-the…) and was very looking forwards to seeing this paper some thoughts 1. bootstrapping learning via ESM2 embeddings (before being fed into another transformer to pretrain) feels increasingly common, I saw it done in the Universal Cell Embeddings paper too. they ablate this and show its useful too, but still feels weird to do… 2. zero shot prediction of binders for new targets is still not possible, despite pretraining a general binding model on 7M~ measurements across 6k targets. for each new target, you need a parent antibody that binds to it + finetuning of mutants of the parent. sad! i wonder how much you need the parent for…would a random set of antibodies for each new target actually be enough? 3. 30k~ new samples (generated via naive mutations to a parent antibody) are required to retrain their general model for new target, but that is within scope to gather in a single run of their internal assay. given that data to finetune their model, the resulting hit rate is quite good, even as edit distance from the parents gets high (by high, i mean 11). take a look at fig 4A 4. their proposed mutations are *across* the entire HCDR1-3 region! not just the regions themselves, but everything in between! decently high diversity on this, which is cool + reasonably new amongst IgG antibody design works (since last i checked) 5. there’s a really interesting section here titled ‘Affinity-Guided Developability Engineering’, showing how their model may be used in practice to do a multi-property-ish optimization thing on top of an antibody with some good characteristics (good binding), but some bad ones (bad developability and high immunogenicity). cool to see how stuff like this could be used in practice to create Actual Drugs overall, cool work. nothing revolutionary, but they dont really claim that it is. slight improvement on some dimensions compared to others, roughly equal on everything else. also one of the better written antibody design papers, most of the others ive read are much harder to understand than this one but the utility of large-scale continuous binding information is less than id hoped. i imagine the desire here is that there is some crazy scaling law to-be-seen, something that requires even more data, but we’ll see how true that ends up being. considering this paper a mild mental update for now final galaxy brained idea: i hope a-alpha bio is looking into mechanistic interpretability! if you’re the only one in the world with a generalized high-diversity binding model, you may be well positioned to learn what **actually** makes for good binders AND good targets (maybe). useful insights to have for a human drug designer probably

I actually do know what to do. It has to be a mass scale economic bill nye type show which is challenging in a fragmented culture but we have to reading rainbowize economics education expeditiously


@MattZeitlin @jaredpolis what's the secret?




Creating the largest protein-protein interaction dataset in the world (A-Alpha Bio) owlposting.com/p/creating-the… i cover @AAlphaBio, an incredibly promising biotech startup ive been evangelizing for over a year now 4.6k words, 22 minutes table of contents in reply! 🧵











