Slavov Laboratory
553 posts


Slavov Laboratory
@slavovLab
We seek principles in the coordination among protein synthesis, metabolism, cell growth and differentiation PI: @slavov_n Videos: https://t.co/Ku19DCIXvX










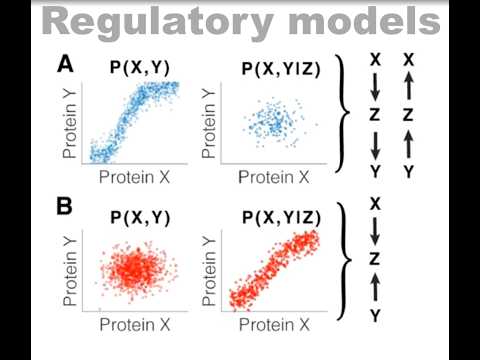

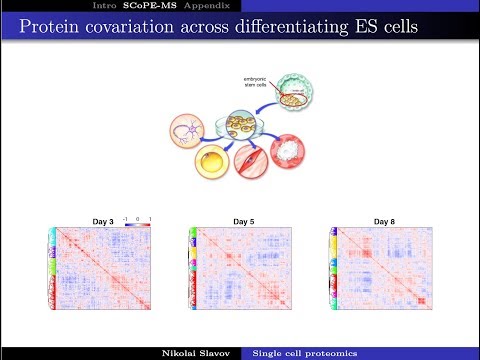





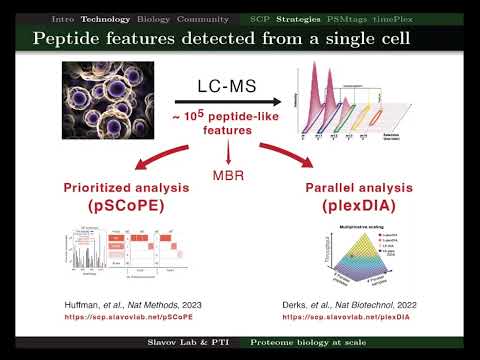

Yesterday I presented a small vignette from two recent preprints at the Cell Size and Growth Seminar. Check out if interested! youtube.com/watch?v=80MPVL…





Protein degradation is often considered a final step in gene expression regulation. It also regulates the first step: Transcription TFs (e.g. p53, HIF, Myc) are regulated by 𝐝𝐞𝐠𝐫𝐚𝐝𝐚𝐭𝐢𝐨𝐧-𝐠𝐚𝐭𝐞𝐝 𝐜𝐨𝐧𝐭𝐫𝐨𝐥: ⬛️ It confers advantages. blog.slavovlab.net/2025/08/16/pro…



Single-Cell Proteomic Technologies: Tools in the Quest for Principles Journal 🔗 annualreviews.org/content/journa… OA 🔗 slavovlab.net/Slavov-Lab-Pub… 2/2