Andrew Leduc
1.9K posts


Andrew Leduc
@_AndrewLeduc
Post-doc @slavovlab Interests: Non-canonical protein sequences Protein degradation Single cell analysis






🐠 Everything we know about biology has been built on an incomplete picture. DNA tells us what a cell might do. Proteins tell us what it’s actually doing. Pumpkinseed announced their $20M Series A today (led by Future Ventures and NfX) to build the platform that reads proteins directly—for the first time. Proteomics has always faced a fundamental constraint: you can only measure what you already know to look for. The current workhorse, mass spectrometry, requires matching protein fragments against reference databases. If a protein isn't in the database, or doesn't ionize reliably, it's invisible. Other approaches rely on fluorescent labels or antibody-based affinity methods, which introduce their own biases and blind spots. The result is a field that has spent decades generating an increasingly detailed map of a small, well-lit corner of the proteome, while biology’s most important data layer remains hidden. This isn't a sensitivity problem. It's a category problem. Existing tools were never designed to read proteins directly de novo. They were designed to find what researchers already suspected was there. Pumpkinseed is built to find everything else. And proteomics is harder than most people outside the field appreciate. When we account for post-translational modifications, non-canonical amino acids, and glycan decorations, there are roughly a thousand distinct chemical monomers in the proteomic alphabet, compared to the four bases of DNA. deSIPHR (de novo Sequencing and Identification of Proteins with High-throughput Raman spectroscopy) is Pumpkinseed's proprietary nanophotonic chip platform, fabricated with semiconducting manufacturing. With over 100 million sensors per square centimeter, it reads proteins, known or unknown, letter by letter — amino acid by amino acid — without a reference catalog of proteins, and at high-throughput. The result is direct, high-resolution proteomic data, including post-translational modifications, non-canonical amino acids, and single-cell detail, that mass spectrometry-based approaches cannot match. What is Raman spectroscopy? Rather than tagging or fragmenting proteins, Raman spectroscopy reads the molecular vibrations of individual molecules. Each amino acid vibrates at a characteristic frequency, producing a unique physical signature that deSIPHR detects directly. This is physics reading biology in the most literal sense. With conventional Raman spectroscopy, only about one in ten million photons interacts with a molecule usefully, far too weak for single-molecule work. Pumpkinseed's answer is a silicon photonic chip patterned with a billion sensors per wafer. Those sensors concentrate light into volumes smaller than a single protein, amplifying Raman scattering efficiency by over 10 million-fold. And their future ventures? “The longer-term ambition is the virtual cell, a computational model that simulates not just how proteins fold but how they interact, respond to drugs, and behave under perturbation inside a living system. AlphaFold demonstrated what structural AI can do once a sequence is known. The gap that cannot be closed is determining the sequence itself from biological samples, particularly for proteins carrying modifications absent from existing databases. Pumpkinseed is designed to supply that input layer. "If the Human Genome Project was the data infrastructure that enabled genomic medicine, we believe the high-resolution proteomic dataset Pumpkinseed is building could be the analogous foundation for AI-driven biological discovery," co-founder Dr. Jen Dionne says. "In our vision, the molecular signatures driving disease, aging, and ecosystem health become fully legible. Medicine shifts from reactive to proactive. Optimal healthspan moves from aspiration to achievable reality." —synbiobeta.com/read/pumpkinse… • The biology mining company: Pumpkinseed.Bio • Today’s News: pumpkinseed.bio/news/pumpkinse…
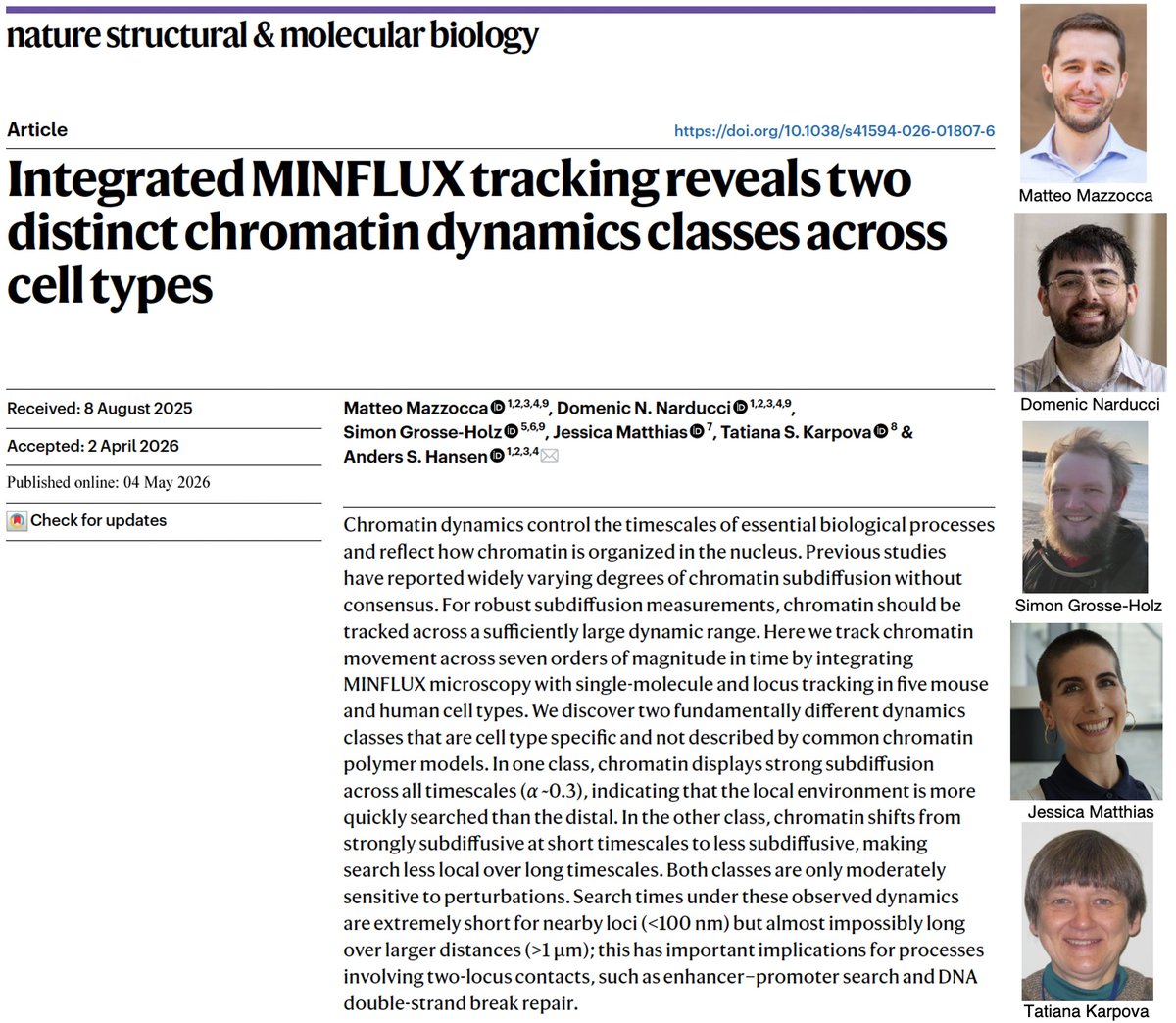
















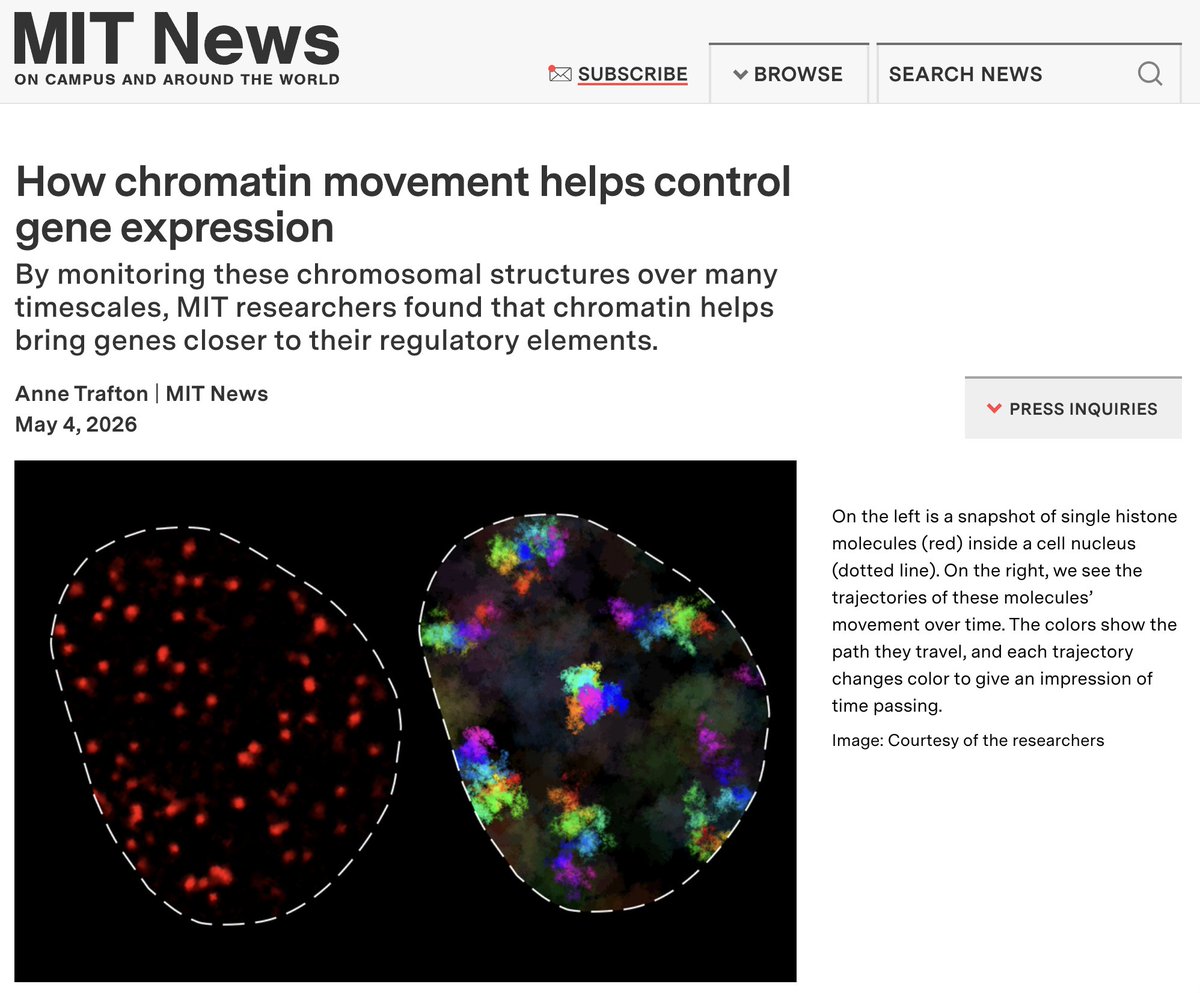
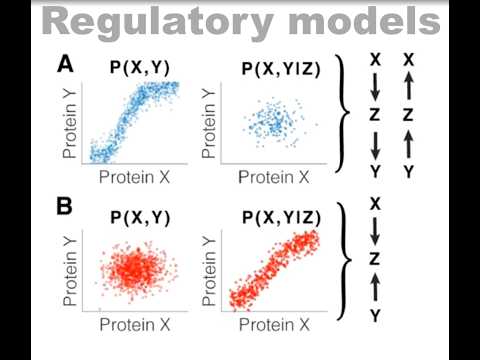
(1/13) Thread on @mazzocca_matteo @DomenicNarducci @SGrosseHolz @_jessematthias new preprint Q: how does chromatin move? Using MINFLUX, SPT & SRLCI, we track chromatin dynamics across 7 orders of magnitude in time to provide some answers biorxiv.org/content/10.110…