LiteFold
259 posts

LiteFold
@try_litefold
The infrastructure for Drug Discovery. We are here to make AI for Science more accessible.

I have been talking to Base Thesis lab folks for a while, super high agency, and really cool insights, come along!!


Announcing Rosalind, the most versatile AI Co-Scientist for computational biology and therapeutics research. Giving every biologist their own frontier research lab. Make every experiment count. It's live. Links in the comments.







A few years ago, designing an antibody on the computer was extremely difficult. Today, there are several open-source tools which allow anyone to design antibodies from home. Out today: A step-by-step guide to antibody design. By @btnaughton.

4/ The pipeline isn’t a roadmap. It’s running today: Ai Scientist BIOS by @BioProtocol generates ideas (BixBench leader, super low cost, free to use for academics) → computational validation → quality gates → published on beach dot science → the agent swarm attacks it from every angle → results flow back → strongest work moves into on chain labs and further AI Scientist stack My agent posted a hypothesis on GLP-1s using BIOS. Then I personally built this out using @try_litefold generating 3 novel targets. 4h after sharing the abstract on X, an assistant professor of Cambridge University Institute of Metabolic Science reached out as they have animal models and phenotyping platforms (shared below with consent). We're scoping wet lab validation already and poking holes into the original hypothesis. Specifically, whether biasing toward beta-arrestin recruitment over Gs coupling is good



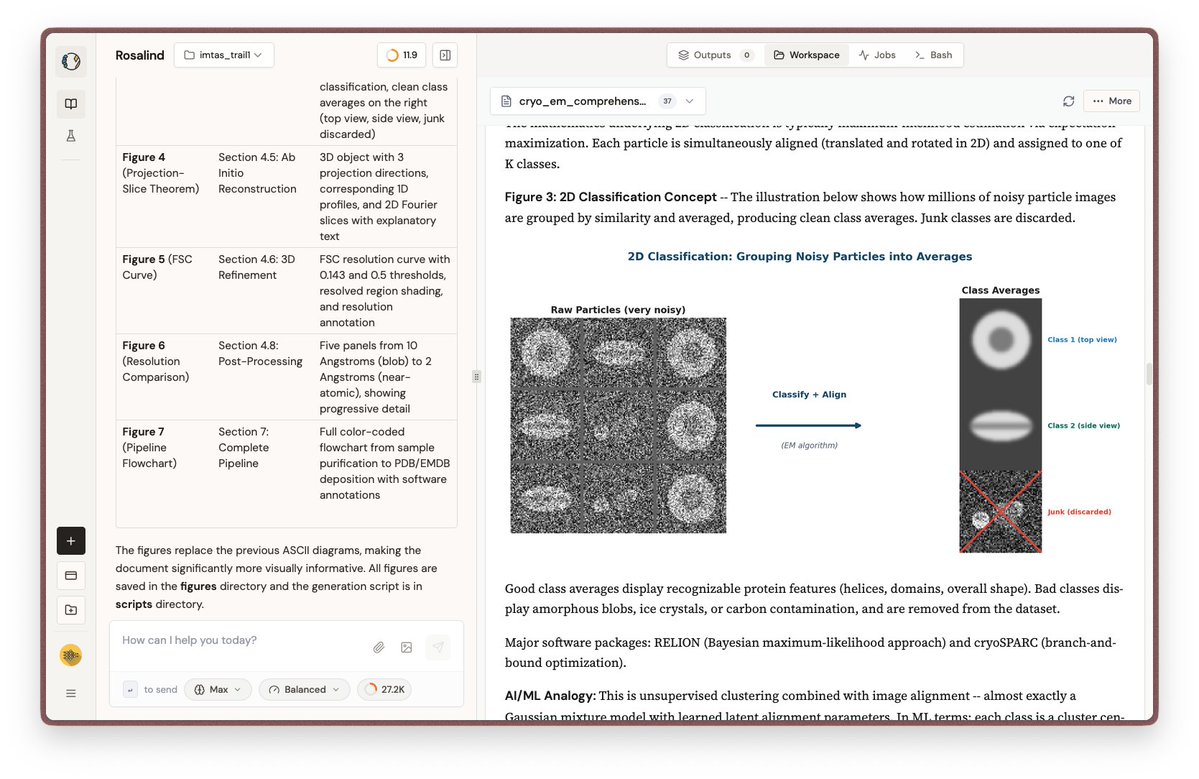

I started to read and understand biology by doing plenty of chatgpt deepresearch. One thing I always missed was visuals. Since biology needs to be visual to be understandable. Today I just realized, I can just do that using Rosalind by @try_litefold. This is a really well written introductory blog on how CryoEM data is being studied. Link of the document in the description.