Sabitlenmiş Tweet

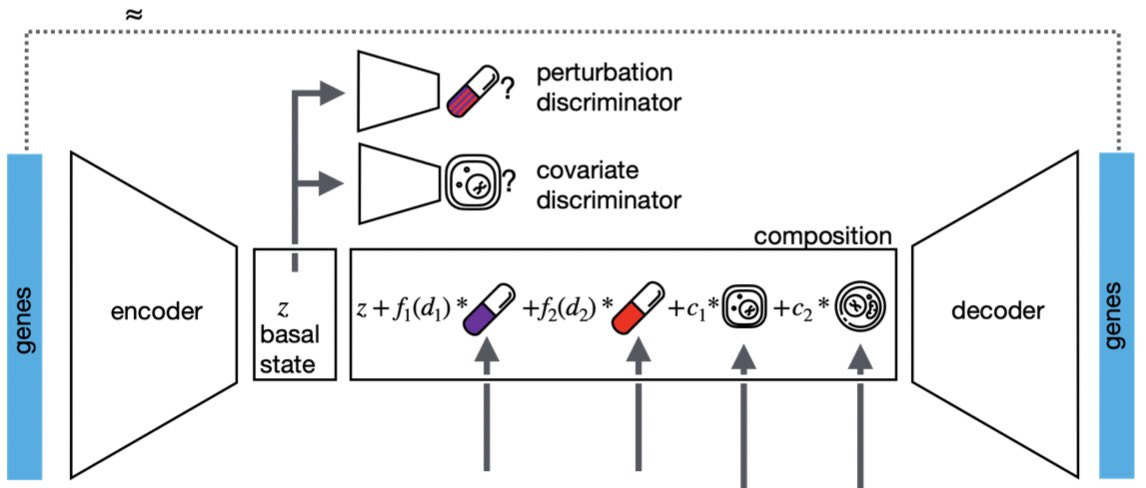
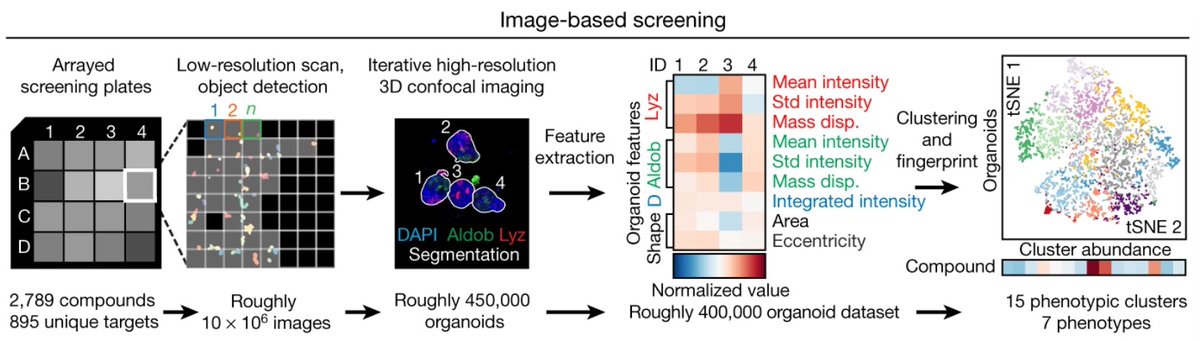
Excited to share Tangram, our method for mapping scRNAseq data on spatial data. Our work is part of the #BICCN and #HubMAP consortia. Collaboration led by Aviv Regev, between @broadinstitute, @Harvard, @skunkworksneu and @unifi.
biorxiv.org/content/10.110…
English