Johannes Hingerl retweetledi

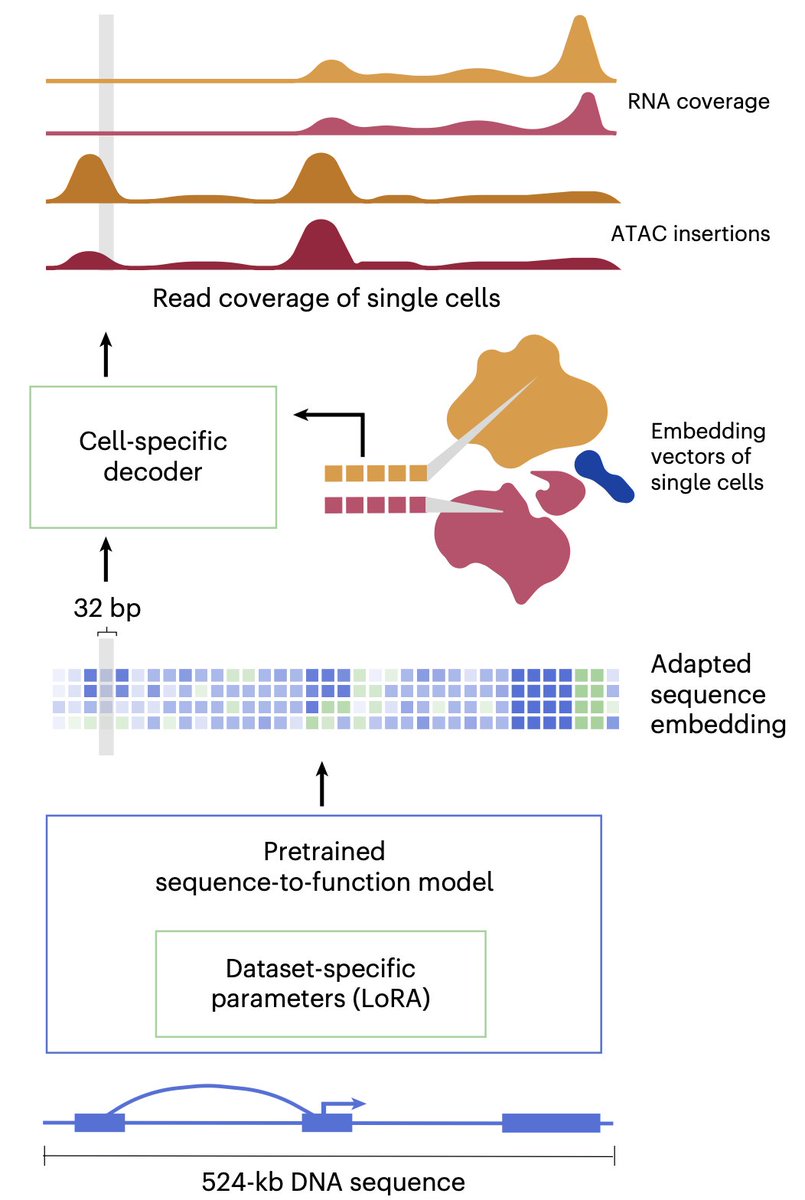
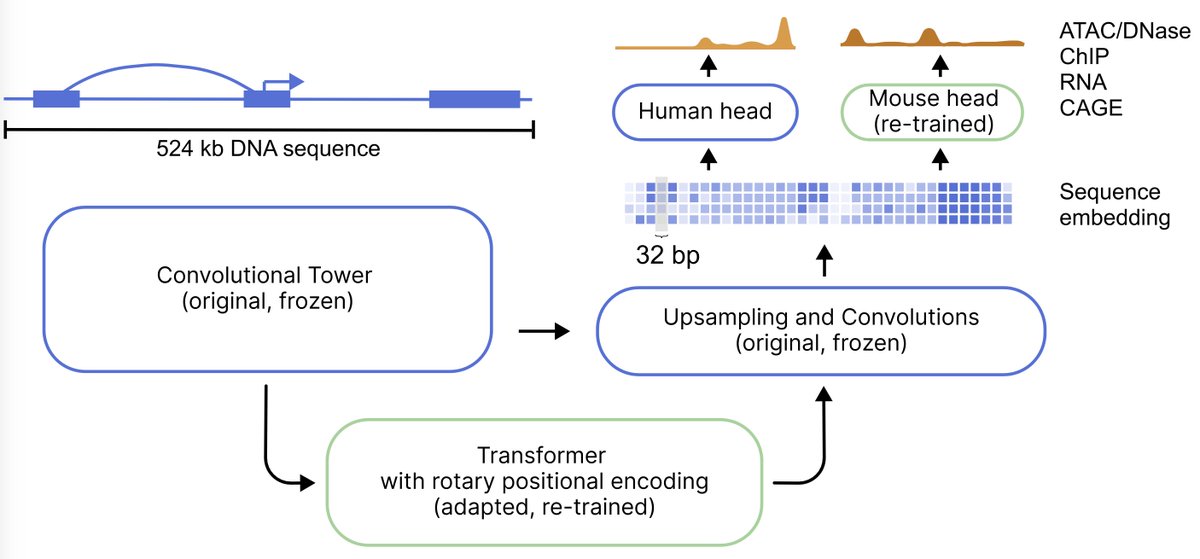
AlphaGenome is out in @nature today along with model weights! 🧬
📄 Paper: nature.com/articles/s4158…
💻 Weights: github.com/google-deepmin…
Getting here wasn’t a straight path. We sat down @googledeepmind to discuss the story behind the model, paper & API: youtu.be/V8lhUqKqzUc

YouTube

English