Read the preprint @ biorxiv.org/content/10.110… and let us know how to improve RADIAnT, send feature requests, get help with problem-solving and more @ github.com/si-ze/RADIAnT
English
Tim Warwick
113 posts


@tim_warwick
Bioinformatics | Chromatin | RNA | Epigenetics Researcher @BrandesLab, @goetheuni, @CPI_ExStra @trr267



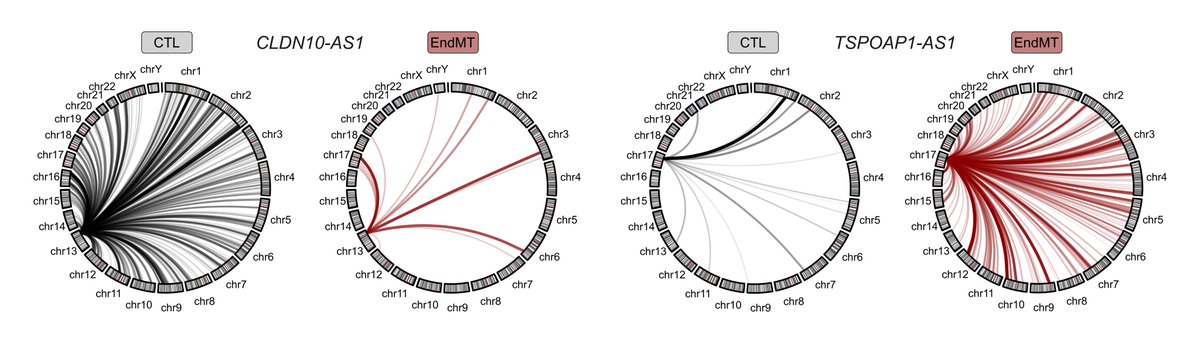

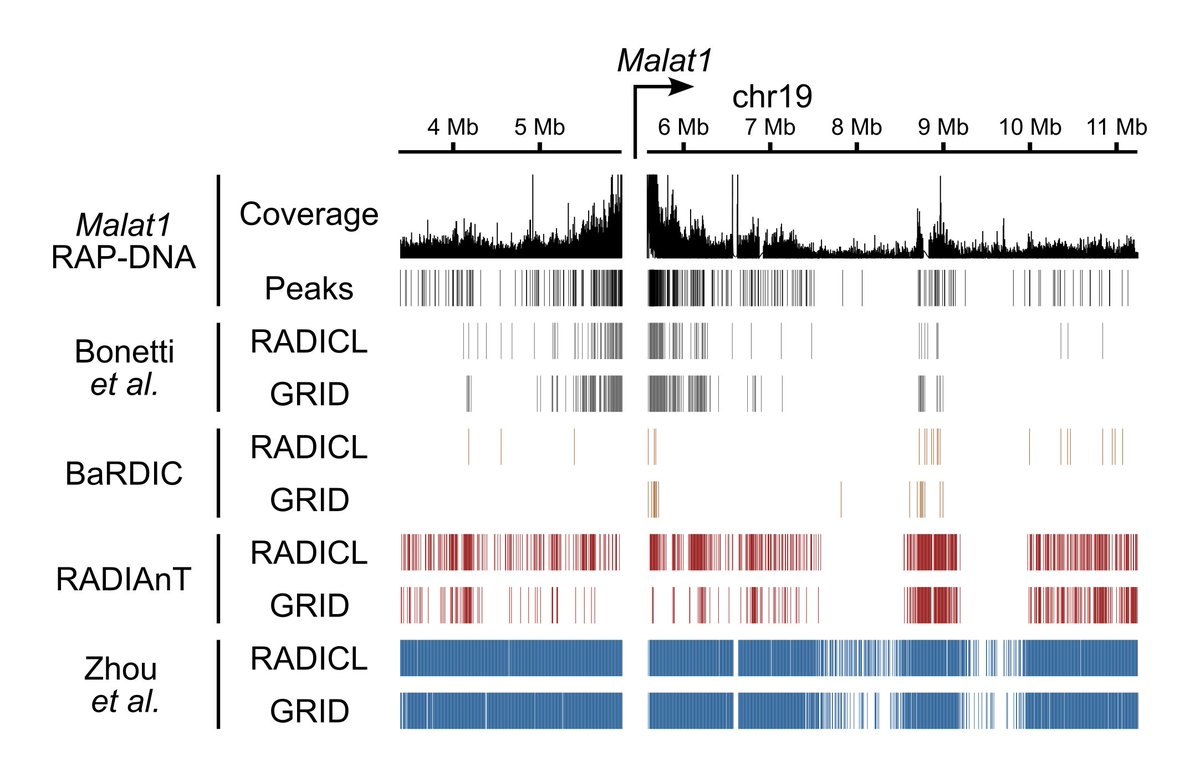









Happy to share our pre-print on #lncRNA recruitment of SWI/SNF to cell type-specific #enhancers. We performed several sequencing experiments to reveal the mechanism for how the highly conserved and ubiquitously expressed SWI/SNF finds its DNA targets genome-wide.