Sabitlenmiş Tweet
Interested in a #postdoc position to develop new sequencing methods? Come work with us! Work includes but is not limited to epigenetics, RNA mods, and single-cell/spatial transcriptomics. docs.google.com/document/d/1ZM…
English
Winston Timp
4.4K posts











Meet Kimberley Billingsley, PhD! 🎉 As a Gates Sr. AD Fellow, she will use high-resolution methylation data to develop predictive models for ADRD, identify epigenetic signatures for better risk stratification and early detection, and establish accurate biomarkers. #ADRD



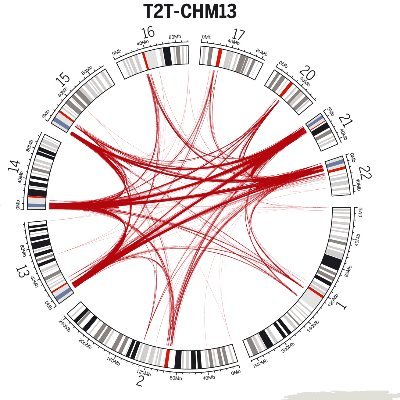
Adaptively integrated sequencing and assembly of near-complete genomes biorxiv.org/content/10.110… #biorxiv_genomic





