Will Spooner
2.3K posts


Will Spooner
@wspoonr
Enabling precision healthcare with big data for genomics





revived my substack because what people think AI will do to drug development timelines vs what it actually can do was driving me crazy. new essay on the two kinds of slow, why the timeline has a floor, and where the real value is: apoorvasrinivasan.substack.com/p/how-much-wil…





Academic co-founder, Peter Campbell, Ph.D., is joining the Quotient team as CSO. We look forward to advancing the first ever #SomaticGenomics platform under his leadership. Read more: bit.ly/3YBCkqS


When writing bioinformatics tools, I often need small datasets to test edge cases or invalid file formats, e.g. files that are truncated, unsorted, have extraneous whitespace, etc. I started compiling examples here: github.com/omgenomics/bio…, contributions are welcome!

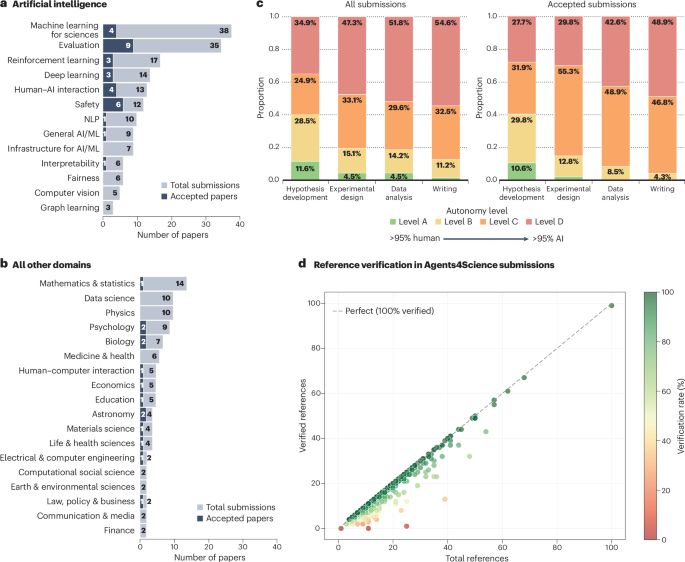
How many academic papers are written with the help of ChatGPT? To answer this question, we analyzed 14mln PubMed abstracts from 2010 to 2024 and looked for excess words: ** Delving into ChatGPT usage in academic writing through excess vocabulary ** arxiv.org/abs/2406.07016 1/11

This never stops being funny: academics trying to write a “lay summary”.




We are Quotient Therapeutics, pioneering #SomaticGenomics to make transformative medicines. Our technology reveals naturally selected genes to target through a variety of therapeutic modalities, enabling near limitless possibilities. quotient-tx.com/news/flagship-…







