The Vaidehi Lab
124 posts


@VaidehiLab
We develop physics-based computational methods to study protein structure, dynamics & function and for drug design | #GPCR #Kinase | *run by the group*










📢👀🔍 We are currently looking for an Associate/Professor level Thoracic Medical Oncologist aa067.taleo.net/careersection/…… 📍City: Los Angeles, CA #lungcancer #thoraciccancer #medicaloncology #ASCO2023 @DrRaviSalgia @JyotiMalhotraMD @ErminiaMassare1 @cityofhope






Mutations in a conserved glutamine residue can confer multiple activation states in alpha subunits of #Gproteins, according to experiments that suggest these proteins do not function as simple on-off switches. @UNC_PHCO @lifescientwists #StructureFunction scim.ag/1Cq





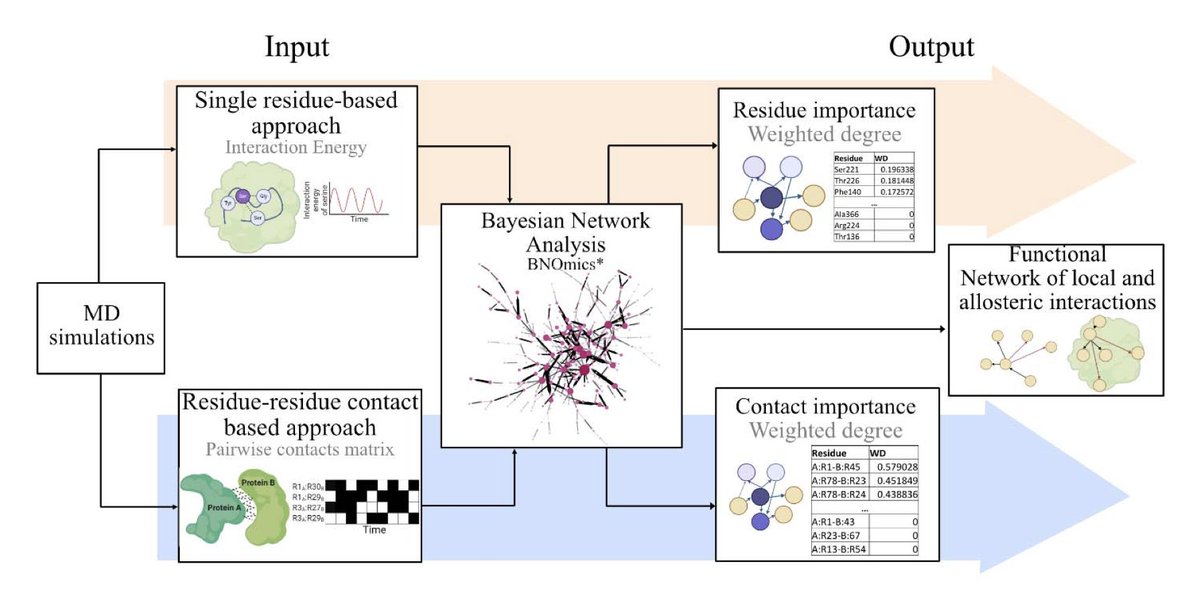
Proud of this work @NatureComms showing persistence of GPCR-G protein interface interactions regulate promiscuity and selectivity of coupling. Impactful contribution by @M5andhu. Thanks to @m_madan_babu and @Laporte001 for great collaboration. nature.com/articles/s4146…


