The countdown to SOT 2026 is on! Visit the 3Brain team from March 22nd – 25th at the San Diego Convention Center.
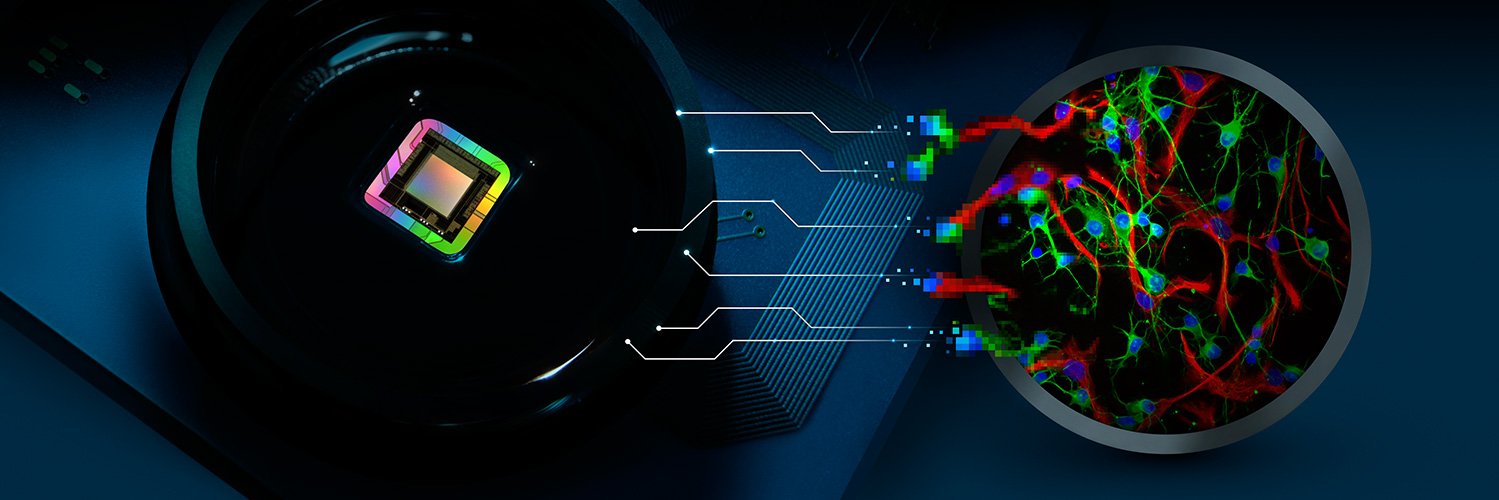
Organoid and spheroid research continues to redefine the boundaries of toxicology, offering unprecedented models for human development, disease and screening purposes. 3Brain is proud to be at the forefront of this shift, we'll be at @SOToxicology to showcase how our CorePlate™ powered HD-MEA platforms can turn complex biological models into actionable data.
Whether you are working with neuronal or cardiac cultures, our range of solutions from the precision of the BioCAM Duplex, to the high-throughput capabilities of the 96-well HyperCAM Delta, our platforms are designed to provide functional insights for enhanced compound screening.
Stop by to explore how our single and multi-well platforms can accelerate your drug screening and neuro/cardiotoxicology research.
#3Brain #HDMEA #Toxicology #Organoids #Neurotoxicology #Electrophysiology #2026SOT
#ToxExpo

English