Martin S. Pedersen
264 posts


Martin S. Pedersen
@Born2bewildtype
Wet lab scientist, into all kinds of microbiology and sequencing. Tweets are my own.


Illumina Complete Long Read Assay yields contiguous bacterial genomes from human gut metagenomes biorxiv.org/cgi/content/sh… #biorxiv_genomic













TurBOT's ready to go! The team at @RegionH are leveraging the fully automated, cutting-edge device to enhance their microbial and clinical research capabilities. Learn more about TurBOT: bit.ly/3wDdL12 More updates coming next week at #nanoporeconf. @Born2bewildtype
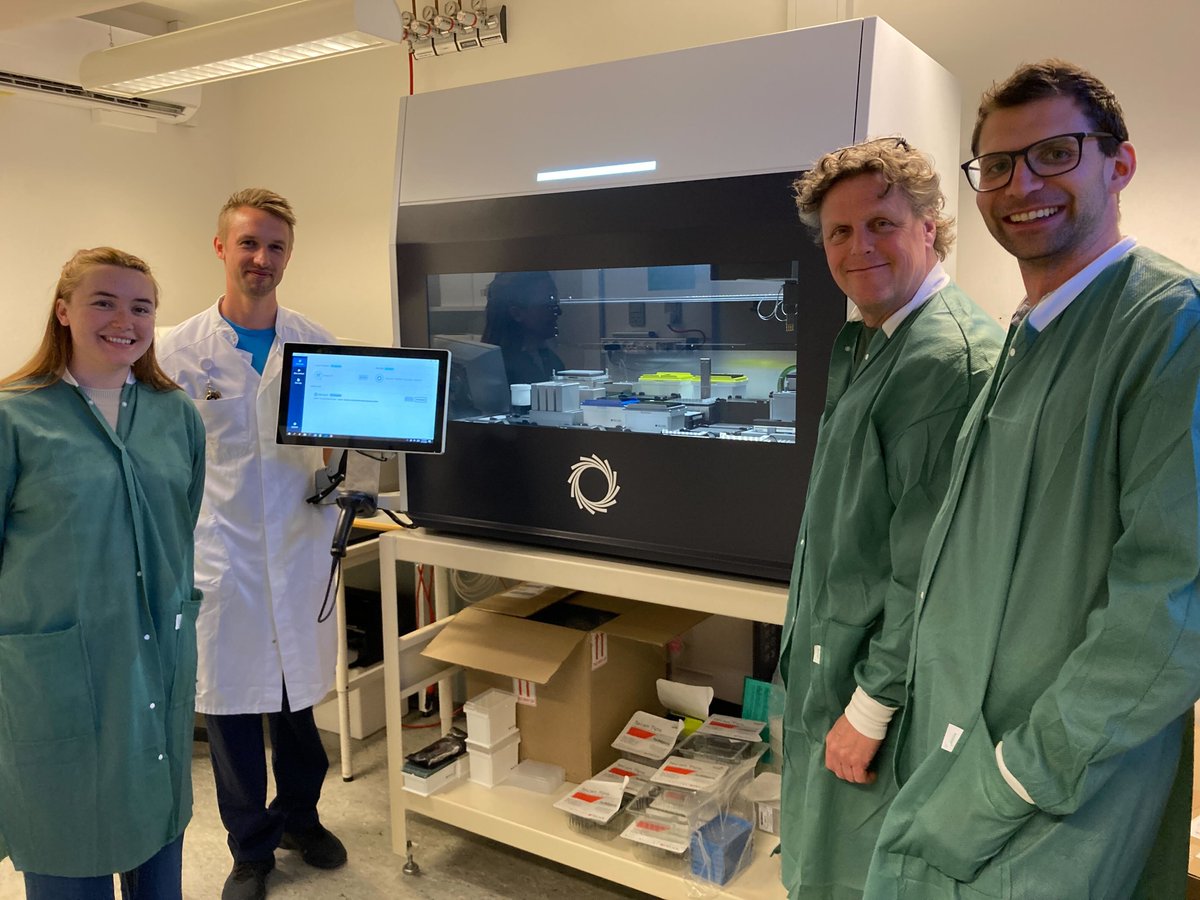











Christmas has come early @nanopore. We are in the process of unpacking the TurBOT, a fully automated, sample-to-sequence solution featuring an integrated MinION sequencer. It is currently in development but you can register your interest on the @nanopore website.



